 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
- The flowchart for construction of PlantTFDB
- Data source
- Data source (TFext)
- Pipeline to construct comprehensive protein dataset
- Family assignment rules
- List of studies which use the Family assignment rules
- Thresholds for domain identification
- Summary of TFs in different taxonomic lineages of green plants and the origination stage of TF families
- Pipeline for parsing BLAST reciprocal best hits (RBHs) and inferring orthologous groups
- Pipeline for GO annotation
- Curation and projection of TF binding motifs
- Transcription factor information
- Multiple sequence alignment
- Phylogenetic trees
- Quick search
- TF prediction server
- Help for PlantRegMap
Family Assignment Rules
After a systematically reviewing
recently-published literatures, we refined family assignment rules. Families that do not meet the criteria ('Transcription factors are
proteins that show sequence-specific DNA binding and are capable of
activating or/and repressing transcription') were excluded in the
updated version, such as transcription cofactors (e.g. AUX/IAA, GIF and
chromatin related proteins like chromatin remodeling factors (e.g.
ARID), histone demethylases (e.g. JUMONJI), histone acetyltransferase
(e.g. PcG ). Moreover, questionable families (e.g. TUBBY-like,
Alfin-like, FHA, and ZIM) were also removed in the updated PlantTFDB. On
the other side, five newly identified TF families (DBB, FAR1, LSD,
NF-X1, STAT) were added. Considering differences in domain composition,
DNA binding specificity and function, AP2/ERF and HB families in
previous PlantTFDB were divided to sub-families. Especially, because it
has been reported that some M type MADS could be potential pseudogenes
or a new class of transposable elements, we also subdivided those M type
MADS to a new subfamily.
In new rules, there were three types of domain
to determine which family a TF belongs to, including DNA-binding domain,
auxiliary domain and forbidden domain. In most cases, DNA-binding domain
self can assign a TF to one certain family correctly. But In some cases,
an auxiliary domain was needed to classify TF into corresponding family.
For example, if a protein only contains a DNA-binding domain - B3, we
can assign it to B3 superfamily. However, B3 superfamily can be divided
into two subfamilies (ARF and B3), and so auxiliary domain need be used
to distinguish these two families: if the protein contains Auxin_resp
domain, it was assigned to ARF family, else it will be assigned to B3
family. Different from previous version, forbidden domains were used to
distinguish proteins containing DNA-binding domain but having no
transcription activity. If a protein contains the forbidden domain, then
we think it was not TF, and should not be collected by PlantTFDB.
Table of family assignment
| Family | DNA-binding domain | Auxiliary domain | Forbidden domain | ||
| AP2/ERF | AP2 | AP2 (>=2) (PF00847) | |||
| ERF | AP2 (1) (PF00847) | ||||
| RAV | AP2 (PF00847)
B3 (PF02362) |
||||
| B3 superfamily | ARF | B3 (PF02362) | Auxin_resp (PF06507) | ||
| B3 | B3 (PF02362) | ||||
| BBR-BPC | GAGA_bind (PF06217) | ||||
| BES1 | DUF822 (PF05687) | ||||
| bHLH | HLH (PF00010) | ||||
| bZIP | bZIP_1 (PF00170) | ||||
| C2C2 | CO-like | zf-B_box (PF00643) | CCT (PF06203) | ||
| Dof | Zf-Dof (PF02701) | ||||
| GATA | GATA-zf (PF00320) | ||||
| LSD | Zf-LSD1 (PF06943) | Peptidase_C14 (PF00656) | |||
| YABBY | YABBY (PF04690) | ||||
| C2H2 | zf-C2H2 (PF00096) | RNase_T (PF00929) | |||
| C3H | Zf-CCCH (PF00642) | RRM_1 (PF00076) or Helicase_C (PF00271) | |||
| CAMTA | CG1 (PF03859) | ||||
| CPP | TCR (PF03638) | ||||
| DBB | zf-B_box (>=2) (PF00643) | ||||
| E2F/DP | E2F_TDP (PF02319) | ||||
| EIL | EIN3 (PF04873) | ||||
| FAR1 | FAR1 (PF03101) | ||||
| GARP | ARR-B | G2-like (self-build) | Response_reg (PF00072) | ||
| G2-like | G2-like (self-build) | ||||
| GeBP | DUF573 (PF04504) | ||||
| GRAS | GRAS (PF03514) | ||||
| GRF | WRC (PF08879) | QLQ (PF08880) | |||
| HB | HD-ZIP | Homeobox (PF00046) | HD-ZIP_I/II (self-build) or SMART (PF01852) | ||
| TALE | Homeobox (PF00046) | BELL (self-build)or ELK (PF03789) | |||
| WOX | homeobox (PF00046) | Wus type homeobox (self-build) | |||
| HB-PHD | homeobox (PF00046) | PHD (PF00628) | |||
| HB-other | homeobox (PF00046) | ||||
| HRT-like | HRT-like (self-build) | ||||
| HSF | HSF_dna_bind (PF00447) | ||||
| LBD (AS2/LOB) | DUF260 (PF03195) | ||||
| LFY | FLO_LFY (PF01698) | ||||
| MADS | M_type | SRF-TF (PF00319) | |||
| MIKC | SRF-TF (PF00319) | K-box (PF01486) | |||
| MYB superfamily | MYB | Myb_dna_bind (>=2) (PF00249) | SWIRM (PF04433) | ||
| MYB_related | Myb_dna_bind (1) (PF00249) | SWIRM (PF04433) | |||
| NAC | NAM (PF02365) | ||||
| NF-X1 | Zf-NF-X1 (PF01422) | ||||
| NF-Y | NF-YA | CBFB_NFYA (PF02045) | |||
| NF-YB | NF-YB (self-build) | ||||
| NF-YC | NF-YC (self-build) | ||||
| Nin-like | RWP-RK (PF02042) | ||||
| NZZ/SPL | NOZZLE (PF08744) | ||||
| S1Fa-like | S1FA (PF04689) | ||||
| SAP | SAP (self-build) | ||||
| SBP | SBP (PF03110) | ||||
| SRS | DUF702 (PF05142) | ||||
| STAT | STAT (self-build) | ||||
| TCP | TCP (PF03634) | ||||
| Trihelix | Trihelix (self-build) | ||||
| VOZ | VOZ (self-build) | ||||
| Whirly | Whirly (PF08536) | ||||
| WRKY | WRKY (PF03106) | ||||
| ZF-HD | ZF-HD_dimer (PF04770) | ||||
- Some proteins containing just one AP2 domain are distinct from proteins from ERF family and more closely related to AP2 family. Those proteins are assignment to AP2 family. Two self-build models are used to distinguish those proteins from ERF proteins.
- Only a subset of C2H2 zinc-finger containing proteins are real TF, many of them maybe not TF and act in RNA metabolism and chromatin-remodeling.
- Some C3H family proteins maybe not TF, some of them regulate RNA stability and some regulate RNA processing.
- ZF-HD_dimer is not DBD but the region involved in the formation of homo and heterodimers. To keep consistent with other rules, we classify it to DBD.
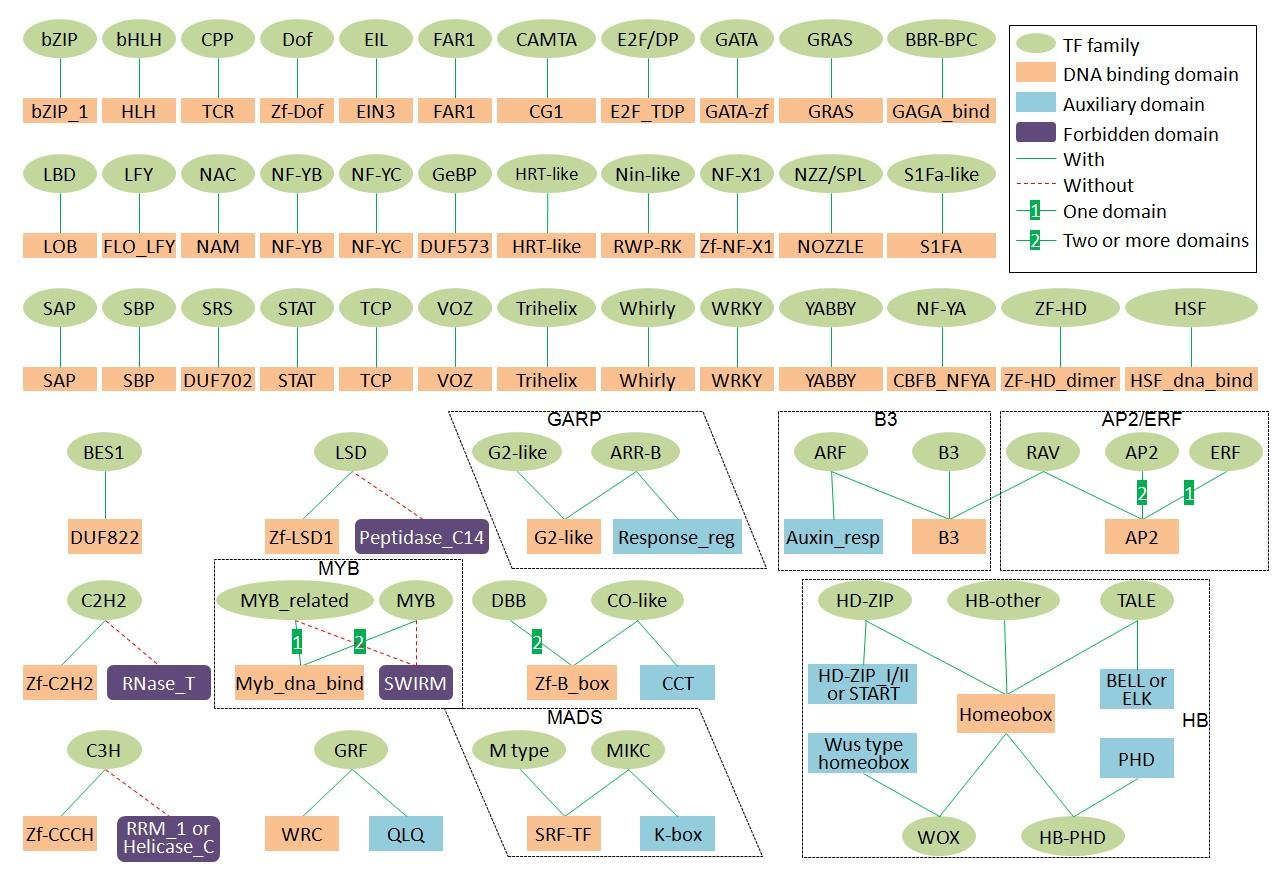
Schema of family assignment
List of studies which use the Family assignment rules
| Species | Common name | Authors | Journal | Year | PMID |
| Ophiorrhiza pumila | - | Rai, A. et al. | Nat Commun | 2021 | 33452249 |
| Syntrichia caninervis | - | Silva, A. T. et al. | Plant J | 2020 | 33277766 |
| Brachypodium distachyon | stiff brome | Kouzai, Y. et al. | Plant J | 2020 | 32891065 |
| Gossypium arboreum | tree cotton | Ashraf, J. et al. | BMC Genomics | 2020 | 32640982 |
| Solanum lycopersicum | tomato | Keller, M. et al. | Sci Rep | 2020 | 32612181 |
| Arachis hypogaea L. | peanut | Zhao, N. et al. | PLoS One | 2020 | 32271855 |
| Chenopodium quinoa | quinoa | Wu, Q. et al. | Plant Physiol Biochem | 2020 | 32289638 |
| Populus trichocarpa | black cottonwood | Chen, H. et al. | Plant Cell | 2019 | 30755461 |
| Oryza sativa L. | rice | Kong, W. et al. | Plants (Basel) | 2019 | 30871082 |
| Medicago truncatula | barrel medic | Sun, L. et al. | Plant J | 2019 | 30776165 |
| Gossypium australe | - | Feng, S. et al. | BMC Plant Biol | 2019 | 31426739 |
| Oryza glaberrima | African rice | Choi, J. Y. et al. | PLoS Genet | 2019 | 30845217 |
| Glycine max | soybean | Gazara, R. K. et al. | Sci Rep | 2019 | 31270425 |
| Trifolium pratense L. | red clover | Chao, Y. et al. | BMC Plant Biol | 2018 | 30477428 |
| Arabidopsis pumila | - | Yang, L. et al. | BMC Genomics | 2018 | 30261913 |
| Arachis ipaensis | - | Lu, Q. et al. | Front Plant Sci | 2018 | 29774047 |
| Santalum album | white sandalwood | Mahesh, H. B. et al. | Plant Physiol | 2018 | 29440596 |
| Foeniculum vulgare Mill. | fennel | Palumbo, F. et al. | Sci Rep | 2018 | 29993007 |



