 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma99g00440.1 | ||||||||
| Common Name | ZOSMA_99G00440 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 400aa MW: 42937.9 Da PI: 7.088 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 121.1 | 1.3e-37 | 67 | 189 | 6 | 114 |
TCP 6 drhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec.eaesssssasnsssg.......... 88
+r++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti WLl+ a+pai+++tgt++++a ++ ++ ++ ++++s
Zosma99g00440.1 67 TRTKDRHTKVEGRGRRIRMPATCAARIFQLTRELGHKSDGETIRWLLEHAEPAIIAATGTGTIPAIAMsVGGTLKVPTQSPSLMassifqgddv 160
89**********************************************************8888855442333333333233333333333333 PP
TCP 89 .....kaaksaakskksqksaasalnlakes 114
k+++s + ++++ a + +n+++e+
Zosma99g00440.1 161 asrkrKKCQS--MPAETKTTAVHPHNYQQEQ 189
2221111111..1111111111112222221 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 9.9E-28 | 68 | 187 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 27.141 | 69 | 123 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 400 aa Download sequence Send to blast |
MEARQKQELR EDDAETDLRM GGGIQDANPL AVFPKPEPVV EERVTPINTV ATLKTGGGRR 60 QAGAVSTRTK DRHTKVEGRG RRIRMPATCA ARIFQLTREL GHKSDGETIR WLLEHAEPAI 120 IAATGTGTIP AIAMSVGGTL KVPTQSPSLM ASSIFQGDDV ASRKRKKCQS MPAETKTTAV 180 HPHNYQQEQS SAAMSMSSGL APIGAVPMWA VSGGRVLSPA LWMLPPPNSA TAATMIPQGG 240 SSTQQQQQQQ QQQQQIFLQQ QQQQIWTAFH HSGSQIINLT ATGPAIPATM FSAVPTLTSL 300 EIPTGSGLQI PMGGFTAEVV AAAGGGGGGQ EPSSFHNHQN RQQKQEHQFM GSSMTSDTSN 360 NYQYQHRVEE DHHDHGDDVS NKEGVQEAVQ PLSQHSPQD* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 162 | 166 | RKRKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00667 | PBM | Transfer from GSVIVG01027588001 | Download |
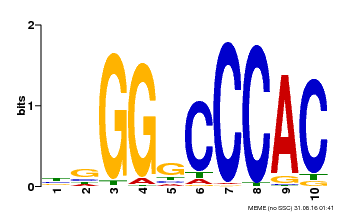
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0K9NHN0 | 0.0 | A0A0K9NHN0_ZOSMR; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3912 | 31 | 67 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45680.1 | 8e-35 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma99g00440.1 |



