 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00003.1.g03070.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 686aa MW: 73683.5 Da PI: 8.9272 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.1 | 8.2e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWT+eEde+l+ +++++G g+W++ ++ g+ R++k+c++rw +yl
Zjn_sc00003.1.g03070.1.am.mk 14 RGRWTAEEDEILASYIAKHGEGSWRSLPKNAGLLRCGKSCRLRWINYL 61
89******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 39.9 | 9.8e-13 | 67 | 173 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT........................................................... CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg........................................................... 22
rg+ ++eE+++++++++ lG+
Zjn_sc00003.1.g03070.1.am.mk 67 RGNISKEEEDIIIKLHATLGNSlkeaqghdsrdprsrlrfgggelssvptpksssrhsvkrtgvgcyvhrwsvpvrvpagl 147
7899****************99*********************************************************** PP
.-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 23 .tWktIartmgkgRtlkqcksrwqkyl 48
tW++Ia++++ gRt++++k++w+++l
Zjn_sc00003.1.g03070.1.am.mk 148 mTWSLIASHLP-GRTDNEIKNYWNSHL 173
***********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-22 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 22.439 | 9 | 65 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.7E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.8E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.57E-20 | 15 | 88 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.22E-11 | 16 | 61 | No hit | No description |
| PROSITE profile | PS50090 | 7.503 | 62 | 173 | IPR017877 | Myb-like domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-20 | 65 | 85 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.1E-11 | 66 | 175 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-20 | 146 | 177 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 3.6E-7 | 149 | 173 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 5.46E-7 | 149 | 178 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.04E-6 | 150 | 173 | No hit | No description |
| Pfam | PF06640 | 8.0E-57 | 191 | 391 | IPR010588 | Myb-related protein P, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0051555 | Biological Process | flavonol biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 686 aa Download sequence Send to blast |
MGRAPCCEKV GLKRGRWTAE EDEILASYIA KHGEGSWRSL PKNAGLLRCG KSCRLRWINY 60 LRADVKRGNI SKEEEDIIIK LHATLGNSLK EAQGHDSRDP RSRLRFGGGE LSSVPTPKSS 120 SRHSVKRTGV GCYVHRWSVP VRVPAGLMTW SLIASHLPGR TDNEIKNYWN SHLSRQIHTY 180 RRIFTAGNDT TITIDISKLH SAGKRRGGRT PGRSALKSTT SSDDKTGSKK VDAEPEPAKT 240 TTKAKETSAF SPAAATSAAS SPPQSDDGAR SVVVDPDQNQ PNSVSDNDGP CSEDASGLMT 300 SLLEEPLDLG LWEAETEMDM EALLLSNTGI DGAAGYSSSF TTGLEAAAGD DLLDMDWDGF 360 AAHLWGDPAQ NNQAAAEPQA AVGCNPEELE SRSDDSGRLR HKDATMETAP PHCEFVIFSY 420 PYQLQRRKYD KRRISLGALR ILAPLAVDAV SCPRTAAGQS PIARHGRGGN GRDDDIMLNQ 480 SIDLHFYRYV FASEILMTRR ATSHPACPAR DRRSADGSVP GGVAKTCLVA LSGELFGPAE 540 CGRRVGWIAH RPAAHLQGTR RAPAEGEDDA ATPPAPDTAR ARQPSASCAL PLRAAAPVPV 600 SSTHPLRFAV TKSGGSAVTE RGNHHRFRLS VGAPVDGRTR ISAWVIIAVV VLPASDPFVS 660 YRARKGGAAQ RLCKGGSYGS SSQANP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
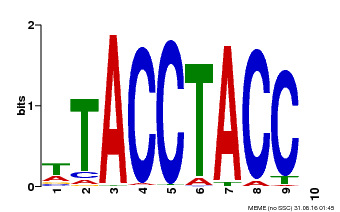
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT039485 | 3e-82 | BT039485.1 Zea mays full-length cDNA clone ZM_BFc0015O24 mRNA, complete cds. | |||
| GenBank | EU959466 | 3e-82 | EU959466.1 Zea mays clone 218238 myb-related protein Myb4 mRNA, complete cds. | |||
| GenBank | HQ858664 | 3e-82 | HQ858664.1 Zea mays clone UT603 MYB transcription factor mRNA, complete cds. | |||
| GenBank | KJ728343 | 3e-82 | KJ728343.1 Zea mays clone pUT6624 MYB-related transcription factor (MYBR102) mRNA, partial cds. | |||
| GenBank | KJ728395 | 3e-82 | KJ728395.1 Zea mays clone pUT6679 MYB transcription factor (MYB3) mRNA, partial cds. | |||
| GenBank | ZMU57002 | 3e-82 | U57002.1 Zea mays P protein (P) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025797897.1 | 1e-115 | myb-related protein P-like | ||||
| TrEMBL | A0A0E0KCP6 | 1e-118 | A0A0E0KCP6_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC03G13890.1 | 1e-119 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G49330.1 | 3e-56 | myb domain protein 111 | ||||



