 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp6g33150 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 336aa MW: 37801.4 Da PI: 5.6238 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 60.5 | 3.5e-19 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ede+lv++v+++G +W++ ++ g++R++k+c++rw +yl
Tp6g33150 14 KGPWTPDEDEKLVNYVQKHGHSSWRALPKLAGLNRCGKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 54.3 | 3.1e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgr++++E++ ++++++ lG++ W+tIa+ ++ gRt++++k++w+++l
Tp6g33150 67 RGRFSPDEEQTILNLHAVLGNK-WSTIANNLP-GRTDNEIKNFWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-25 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.363 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.8E-31 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.6E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.05E-12 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.146 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-27 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.5E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.6E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.11E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MGRSPCSDDS GLKKGPWTPD EDEKLVNYVQ KHGHSSWRAL PKLAGLNRCG KSCRLRWTNY 60 LRPDIKRGRF SPDEEQTILN LHAVLGNKWS TIANNLPGRT DNEIKNFWNT HLKKKLIQMG 120 IDPMTHRPRT DIFSGLSQLM SLSSNLRGFV DLQQQFPIDQ EQTILKLQTE MAKFQLFQCL 180 LQPPSMNNNI NPNDFDTLSL LNSIASFKDT TSNNLDLGTY LQDFNSLPSL KTLNSNIGPS 240 SVFPQDPNDD HFKFFTQREN LPVSPIWLSD PSSSNQSMLP ALDPSSALSD EMIRNQYVIE 300 DVNSNLTSSS CQESGASASA AWPDHLLDDS IFSDIP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-29 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250|UniProtKB:Q9M2D9}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00503 | DAP | Transfer from AT5G10280 | Download |
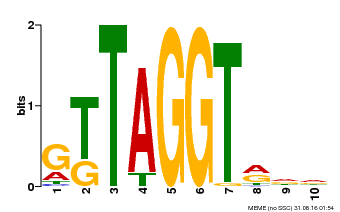
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp6g33150 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by gravity in roots. {ECO:0000269|PubMed:24902892}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM975660 | 0.0 | KM975660.1 Brassica napus MYB transcription factor 92.1 (MYB92.1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006399522.1 | 0.0 | transcription factor MYB92 | ||||
| Swissprot | Q9SBF3 | 0.0 | MYB92_ARATH; Transcription factor MYB92 | ||||
| TrEMBL | A0A1J3GG69 | 0.0 | A0A1J3GG69_NOCCA; Transcription factor MYB39 | ||||
| STRING | XP_006399522.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10280.1 | 0.0 | myb domain protein 92 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



