 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G10280.1 | ||||||||
| Common Name | ATMYB64, ATMYB92, F18D22_50, MYB92 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 334aa MW: 37653 Da PI: 6.1765 | ||||||||
| Description | myb domain protein 92 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 60.5 | 3.5e-19 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ede+lv++v+++G +W++ ++ g++R++k+c++rw +yl
AT5G10280.1 14 KGPWTPDEDEKLVNYVQKHGHSSWRALPKLAGLNRCGKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.8 | 4.4e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgr++++E++ +++++ lG++ W+tIa+ ++ gRt++++k++w+++l
AT5G10280.1 67 RGRFSPDEEQTILNLHSVLGNK-WSTIANQLP-GRTDNEIKNFWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-24 | 7 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.363 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.27E-31 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.6E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.00E-12 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.031 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-27 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.3E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.53E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000981 | Molecular Function | RNA polymerase II transcription factor activity, sequence-specific DNA binding | ||||
| GO:0001135 | Molecular Function | transcription factor activity, RNA polymerase II transcription factor recruiting | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MGRSPISDDS GLKKGPWTPD EDEKLVNYVQ KHGHSSWRAL PKLAGLNRCG KSCRLRWTNY 60 LRPDIKRGRF SPDEEQTILN LHSVLGNKWS TIANQLPGRT DNEIKNFWNT HLKKKLIQMG 120 FDPMTHRPRT DIFSGLSQLM SLSSNLRGFV DLQQQFPIDQ EHTILKLQTE MAKLQLFQYL 180 LQPSSMSNNV NPNDFDTLSL LNSIASFKET SNNTTSNNLD LGFLGSYLQD FHSLPSLKTL 240 NSNMEPSSVF PQNLDDNHFK FSTQRENLPV SPIWLSDPSS TTPAHVNDDL IFNQYGIEDV 300 NSNITSSSGQ ESGASASAAW PDHLLDDSIF SDIP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-29 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.5999 | 0.0 | root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30683090 | 0.0 | ||||
| Genevisible | 250450_at | 0.0 | ||||
| Expression Atlas | AT5G10280 | - | ||||
| AtGenExpress | AT5G10280 | - | ||||
| ATTED-II | AT5G10280 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in roots and at lower levels in stems, flowers and siliques. {ECO:0000269|PubMed:24902892}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a putative transcription factor (MYB92). | |||||
| UniProt | Probable transcription factor. {ECO:0000250|UniProtKB:Q9M2D9}. | |||||
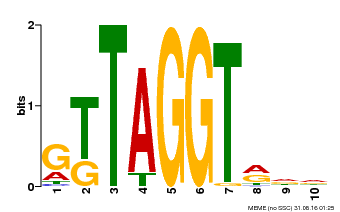
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00503 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G10280.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by gravity in roots. {ECO:0000269|PubMed:24902892}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | jasmonic acid, salicylic acid | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G10280 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY519619 | 0.0 | AY519619.1 Arabidopsis thaliana MYB transcription factor (At5g10280) mRNA, complete cds. | |||
| GenBank | BT002877 | 0.0 | BT002877.1 Arabidopsis thaliana clone RAFL14-88-D17 (R20400) putative myb family transcription factor (At5g10280) mRNA, complete cds. | |||
| GenBank | BT004402 | 0.0 | BT004402.1 Arabidopsis thaliana clone U20400 putative myb family transcription factor (At5g10280) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_196590.1 | 0.0 | myb domain protein 92 | ||||
| Swissprot | Q9SBF3 | 0.0 | MYB92_ARATH; Transcription factor MYB92 | ||||
| TrEMBL | A0A178UPI6 | 0.0 | A0A178UPI6_ARATH; MYB92 | ||||
| STRING | AT5G10280.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 | Representative plant | OGRP5 | 17 | 1784 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G10280.1 |
| Entrez Gene | 830892 |
| iHOP | AT5G10280 |
| wikigenes | AT5G10280 |



