 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp2g26380 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 440aa MW: 48826.1 Da PI: 6.8189 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 43.9 | 4e-14 | 107 | 129 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
e C+f++rtG CkyG++CkF+H+
Tp2g26380 107 EDCSFYMRTGSCKYGSSCKFNHP 129
89********************9 PP
| |||||||
| 2 | zf-CCCH | 29.8 | 9.9e-10 | 155 | 175 | 5 | 25 |
--SGGGGTS--TTTTT-SS-S CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaH 25
+C++++rtG CkyG++C+F+H
Tp2g26380 155 ECKYYFRTGGCKYGETCRFSH 175
6******************** PP
| |||||||
| 3 | zf-CCCH | 37.5 | 3.9e-12 | 198 | 222 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
++++C+f++r+G Ck+G+ CkF+H+
Tp2g26380 198 GEKECPFYMRNGSCKFGGDCKFNHP 222
7899********************9 PP
| |||||||
| 4 | zf-CCCH | 18.4 | 4e-06 | 341 | 366 | 2 | 27 |
-S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHgp 27
++++C+++ +tG Ck+ +Ck++H++
Tp2g26380 341 DQPECSYYVKTGDCKFKYKCKYHHPK 366
689*********************96 PP
| |||||||
| 5 | zf-CCCH | 32.1 | 1.9e-10 | 389 | 411 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
C +++r+G+Ck+G+ C+F H+
Tp2g26380 389 NMCTHYSRYGICKFGPACRFDHS 411
67********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF90229 | 1.28E-6 | 101 | 131 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.159 | 103 | 131 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.4E-6 | 103 | 130 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.4E-11 | 107 | 129 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 6.5E-6 | 109 | 128 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.699 | 150 | 178 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.0E-4 | 150 | 177 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.44E-6 | 151 | 179 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 3.2E-7 | 155 | 175 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.0E-5 | 156 | 178 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.601 | 196 | 224 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 2.8E-6 | 196 | 223 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 8.24E-6 | 198 | 223 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 9.4E-10 | 198 | 222 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.577 | 339 | 367 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.076 | 339 | 366 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 9.16E-5 | 343 | 369 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.971 | 385 | 413 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.0078 | 385 | 412 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.85E-6 | 386 | 413 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.4E-4 | 390 | 411 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 5.1E-7 | 391 | 411 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 440 aa Download sequence Send to blast |
MSKPEEFPDP NPAIPDPKPP SRSSSDNDPA ADRAPSDLNH ISEELTDNLR NVSLDENTKE 60 VTGVPISVSE GNGETGSKEG SEEGGGRESE STVEKRMIVY PVRPDAEDCS FYMRTGSCKY 120 GSSCKFNHPV RRKLQIGREK VREREADVEN PRLMECKYYF RTGGCKYGET CRFSHTKEQT 180 SLASRPELNF LGLPIRPGEK ECPFYMRNGS CKFGGDCKFN HPDPTAIGGV DSPLFRGTNG 240 GSVGSFSPKD APQASSTSWS SSRHMNGTGT APFIPVMYSQ NRGVSPQTPE WNGYQAPSVY 300 PPERSVLPPS TYPANNSLGE TSSFSQFQPQ MPVEEFPERP DQPECSYYVK TGDCKFKYKC 360 KYHHPKNRLP KQIPSSFNDK GLPLRPDQNM CTHYSRYGIC KFGPACRFDH SIPPTFSSSS 420 SQTVEVPQVG GKGDENDSWH |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
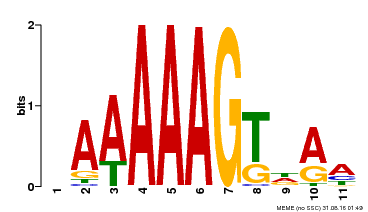
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp2g26380 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394295.1 | 0.0 | zinc finger CCCH domain-containing protein 67 isoform X1 | ||||
| Swissprot | Q5RJC5 | 0.0 | C3H67_ARATH; Zinc finger CCCH domain-containing protein 67 | ||||
| TrEMBL | V4MJP0 | 0.0 | V4MJP0_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006394295.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4176 | 27 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.1 | 0.0 | C3H family protein | ||||



