 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG013400t2 | ||||||||
| Common Name | TCM_013400 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 467aa MW: 51702.6 Da PI: 6.5157 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 26.8 | 1.2e-08 | 59 | 84 | 1 | 27 |
TSSS-HHHHHHHHHHHHHTTTT-HHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktI 27
r +WT+eE++++++a +++G g W+ I
Thecc1EG013400t2 59 REKWTEEEHQKFLEALRLYGRG-WRQI 84
789*******************.*998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.18E-7 | 48 | 85 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-5 | 59 | 86 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.9E-6 | 59 | 84 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.26E-4 | 61 | 84 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MATQDQVEGT TSPNALKSGI CCSNSSPQCE TLTQFQELYT FKHDHTPKVR KPYTITKQRE 60 KWTEEEHQKF LEALRLYGRG WRQIEGFQVV RESNGGFEGS INPIEIPPPR PKRKPVHPYP 120 RKSVDSLKGI SPSSEPERSP SPSQFVREQD NKSPTSVLSA LTSDAMGSAA SEQQNGCSSP 180 TSCTTNMQSI NTSPVEKDID YATSNSSAEE EKASLSSVKV FGHSAVEDVL PMKLNADFKG 240 SVGAKGDAKM VVPFTSIKLF GKTVQVKDSR KPSMDAENFK SPTSKTAQGD IDAEGDMLVQ 300 ALPSTHLDTR LSLGTVNEDW SVVPSQANLS PYMEIHPDKL DHVESTSDAP LPWWTFYQGL 360 PFYYITSFNQ TQTDSCVEER VKQKEILNER SSTGSNTGSV SQAENREKSS YSVDSQCQRP 420 CPEGKTTLQK CSRGFVPYKR CLAERDMSSS VVMSEERERQ RSRVCS* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
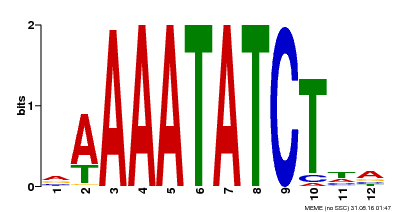
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007037069.1 | 0.0 | PREDICTED: protein REVEILLE 7 | ||||
| TrEMBL | A0A061FWD0 | 0.0 | A0A061FWD0_THECC; Homeodomain-like superfamily protein, putative isoform 2 | ||||
| STRING | EOY21570 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 8e-26 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG013400t2 |
| Entrez Gene | 18604492 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



