 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G18330.2 | ||||||||
| Common Name | EPR1, F15H18.16, RVE7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 372aa MW: 41541.6 Da PI: 8.9054 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.5e-17 | 76 | 120 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +W++eE++++++a+k++G g W+ I +++g ++t+ q++s+ qk+
AT1G18330.2 76 REKWSEEEHDRFLEAIKLYGRG-WRQIQEHIG-TKTAVQIRSHAQKF 120
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.4E-16 | 70 | 124 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.768 | 71 | 125 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-9 | 72 | 123 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.5E-16 | 74 | 123 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.2E-13 | 75 | 123 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.4E-15 | 76 | 119 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.82E-11 | 78 | 121 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MLCFVRFQAG FVRIIVAARK RFRYFLMAAE DRSEELSSNV ENGSCNSNEG INPETSSHWI 60 ENVVKVRKPY TVTKQREKWS EEEHDRFLEA IKLYGRGWRQ IQEHIGTKTA VQIRSHAQKF 120 FSKMAQEADS RSEGSVKAIV IPPPRPKRKP AHPYPRKSPV PYTQSPPPNL SAMEKGTKSP 180 TSVLSSFGSE DQVNRCSSPN SCTSDIQSIG ATSIDKKNNY TTSKQPFKDD SDIGSTPISS 240 ITLFGKIVLV AEESHKPSSY NDDDLKQMTC QENHYSGMLV DTNLSLGVWE TFCTGSNAFG 300 SVTEASENLE KSAEPISSSW KRLSSLEKQG SCNPVNASGF RPYKRCLSER EVTSSLTLVA 360 SDEKKSQRAR IC |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.27131 | 0.0 | flower| silique| vegetative tissue | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 261663_at | 0.0 | ||||
| Expression Atlas | AT1G18330 | - | ||||
| AtGenExpress | AT1G18330 | - | ||||
| ATTED-II | AT1G18330 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | EARLY-PHYTOCHROME-RESPONSIVE1 | |||||
| UniProt | Transcription factor involved in phytochrome A-mediated cotyledon opening. Controlled by the central oscillator mediated by LHY and CCA1. Part of a regulatory circadian feedback loop. Regulates its own expression. {ECO:0000269|PubMed:14523250}. | |||||
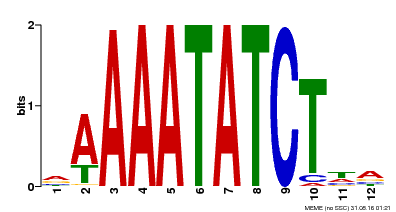
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G18330.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of transcript abundance near subjective dawn. Up-regulated transiently by red light and far red light. {ECO:0000269|PubMed:14523250, ECO:0000269|PubMed:19805390}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Upstream Regulators) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator (A: Activate/R: Repress) | |||||
| ATRM | AT5G10140 (R) | |||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | ethylene | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: No effect on the regulation of core clock associated genes or on the hypocotyl length. Rve1, rve2 and rve7 triple mutant has no alteration in the period or phase of the clock. {ECO:0000269|PubMed:19805390}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G18330 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB115696 | 0.0 | AB115696.1 Arabidopsis thaliana EPR1 mRNA for MYB-related transcription factor EPR1, complete cds. | |||
| GenBank | AY063952 | 0.0 | AY063952.1 Arabidopsis thaliana unknown protein (At1g18330) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001117304.1 | 0.0 | Homeodomain-like superfamily protein | ||||
| Swissprot | B3H5A8 | 0.0 | RVE7_ARATH; Protein REVEILLE 7 | ||||
| TrEMBL | D7KGC9 | 0.0 | D7KGC9_ARALL; Early-phytochrome-responsive1 | ||||
| STRING | AT1G18330.2 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9174 | 27 | 38 | Representative plant | OGRP1255 | 15 | 49 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G18330.2 |
| Entrez Gene | 838413 |
| iHOP | AT1G18330 |
| wikigenes | AT1G18330 |



