 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo3G0014200 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | WOX | ||||||||
| Protein Properties | Length: 263aa MW: 28470.8 Da PI: 8.2505 | ||||||||
| Description | WOX family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 48.2 | 1.9e-15 | 28 | 88 | 2 | 57 |
T--SS--HHHHHHHHHHHHH.SSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 2 rkRttftkeqleeLeelFek.nrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
r+R+t+++eq+ +Le+ F++ +p +++ ++ k l ++ +++V++WFqNrR + ++
Spipo3G0014200 28 RTRWTPKPEQILILESIFNSgMVNPPKDDTVRIRKLLerfgSVGDANVFYWFQNRRSRSRR 88
89*****************99*************************************997 PP
| |||||||
| 2 | Wus_type_Homeobox | 105.8 | 2.7e-34 | 27 | 90 | 2 | 65 |
Wus_type_Homeobox 2 artRWtPtpeQikiLeelyksGlrtPnkeeiqritaeLeeyGkiedkNVfyWFQNrkaRerqkq 65
+rtRWtP+peQi iLe++++sG+++P+k+ + ri++ Le++G ++d+NVfyWFQNr++R+r++q
Spipo3G0014200 27 VRTRWTPKPEQILILESIFNSGMVNPPKDDTVRIRKLLERFGSVGDANVFYWFQNRRSRSRRRQ 90
589***********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-8 | 8 | 89 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.01E-13 | 24 | 94 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 11.418 | 24 | 89 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.0E-5 | 26 | 93 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 5.1E-13 | 28 | 88 | IPR001356 | Homeobox domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048830 | Biological Process | adventitious root development | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MEDQARKAKS RPHGSAGDVV FPAAERVRTR WTPKPEQILI LESIFNSGMV NPPKDDTVRI 60 RKLLERFGSV GDANVFYWFQ NRRSRSRRRQ RQLQAGIAAE PRALSATDGG GGGGAARCDV 120 TLPGAFANSS TNSSFGAFVP SSTSDVSIGT NFSPDIFADD SAEDLFSISR QMGFTETIDQ 180 SRRYAPPPAR HHGASSFHYQ SGLITVFING IPQEVPSGPF DTKKVFGQDA MLLHSSGELV 240 PMNDCGTLFS ALHNGESYIL LS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in crown root development (PubMed:26307379, PubMed:29740464). Promotes the development of crown roots (both initiation and elongation), main components of the fibrous root system (PubMed:26307379, PubMed:29740464). Regulates the expression of genes required for crown root development, and hormone-responsive genes involved in cytokinin signaling (e.g. RR1, RR2, RR3 and RR4), and auxin signaling (e.g. IAA5, IAA11, IAA23 and IAA31) (PubMed:26307379). Functions dowstream of the auxin biosynthetic genes YUCCA1 in the promotion of crown root development (PubMed:29740464). {ECO:0000269|PubMed:26307379, ECO:0000269|PubMed:29740464}. | |||||
| UniProt | Transcription factor which may be involved in developmental processes. Promotes the development of crown roots (both initiation and elongation), main components of the fibrous root system, by regulating the expression of genes required for crown root development and hormone-responsive genes involved in cytokinin (e.g. RR1, RR2, RR3 and RR4) and auxin (e.g. IAA5, IAA11, IAA23 and IAA31) signaling. {ECO:0000250|UniProtKB:Q0D3I7}. | |||||
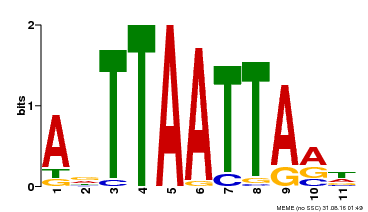
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00330 | DAP | Transfer from AT3G03660 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022719652.1 | 2e-73 | WUSCHEL-related homeobox 11-like | ||||
| Swissprot | B8B644 | 5e-57 | WOX11_ORYSI; WUSCHEL-related homeobox 11 | ||||
| Swissprot | Q0D3I7 | 5e-57 | WOX11_ORYSJ; WUSCHEL-related homeobox 11 | ||||
| TrEMBL | A0A1D1XY95 | 8e-81 | A0A1D1XY95_9ARAE; WUSCHEL-related homeobox 11 (Fragment) | ||||
| STRING | XP_008781307.1 | 1e-71 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1078 | 37 | 109 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G03660.1 | 6e-53 | WUSCHEL related homeobox 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo3G0014200 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



