 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo26G0027000 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 620aa MW: 68495.5 Da PI: 6.8412 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.8 | 3.9e-13 | 465 | 511 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI++Lq
Spipo26G0027000 465 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAISYINDLQ 511
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.9E-51 | 52 | 239 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.282 | 461 | 510 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.33E-15 | 464 | 515 | No hit | No description |
| SuperFamily | SSF47459 | 6.15E-19 | 464 | 524 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-10 | 465 | 511 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.3E-18 | 465 | 524 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.5E-17 | 467 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 620 aa Download sequence Send to blast |
MKTEKGTVGV GFWEEEDKAM AVAVLGAKAF DYITTVHVSS EGLTAIGGDS DLQEKLVALV 60 DGPNPSGFTW NYAIFWQISR SATGDIVLGW GDGHCREPRQ REDSSQRGWQ QHLDEAHQHM 120 RKQVLQKLSV FAGGSDDENY ALRLDRVTDT EMFFLASMYF SFPRGEGAPG KVFASGNHLW 180 ISDALPNSSL SDFCVRGFLA RSTGMRTIVL VPTTNGVLEL GSVAPVSENA EAMQTIKSIL 240 SSGPAKPVPG GSDRGDSQIT AANASSFGFM PRGEERPRIF GKDLNLGQPQ TDERLSVAKM 300 EEQRQQWDLE SSNGGGGDRI RSPDSRKDLH LLNWTQSHRV GANQAHVNNQ QKFSSGVVIT 360 EADACRRPFG HYSNGAREDP QQSHFPPKRQ RSLPAPLTQP RQIDFSRAAA TSMAVPVESQ 420 PNSLQAEQSG ANLASSKEDR PFPPEERRPR KRGRKPANGR EEPLNHVEAE RQRREKLNQR 480 FYALRAVVPN ISKMDKASLL GDAISYINDL QRKLKEMESE RERFSDVTVV DARKRVECPF 540 VDVQVVHNEV VVHVSCPLET HPVSKVISVF NEEQIDVVDS KISAGSDTVF HTFVVRPQGS 600 EQLTKDRLVA ALSREANTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 2e-27 | 459 | 521 | 2 | 64 | Transcription factor MYC2 |
| 5t0f_A | 4e-26 | 52 | 236 | 9 | 188 | Transcription factor MYC3 |
| 5t0q_A | 4e-26 | 52 | 236 | 9 | 188 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 446 | 454 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
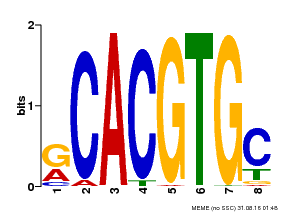
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010242396.1 | 0.0 | PREDICTED: transcription factor bHLH13-like | ||||
| Refseq | XP_010242398.1 | 0.0 | PREDICTED: transcription factor bHLH13-like | ||||
| Refseq | XP_010242399.1 | 0.0 | PREDICTED: transcription factor bHLH13-like | ||||
| Refseq | XP_019051449.1 | 0.0 | PREDICTED: transcription factor bHLH13-like | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A1D1ZHC5 | 0.0 | A0A1D1ZHC5_9ARAE; Transcription factor bHLH13 | ||||
| STRING | XP_010242395.1 | 0.0 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-137 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo26G0027000 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



