 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G01260.3 | ||||||||
| Common Name | BHLH13, EN39, F6F3.7, JAM2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 590aa MW: 66002 Da PI: 6.7082 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.9 | 7.5e-13 | 433 | 478 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L Av YI++L
AT1G01260.3 433 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAVSYINEL 478
799***********************66......***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 7.1E-53 | 48 | 236 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.326 | 429 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.17E-13 | 432 | 483 | No hit | No description |
| SuperFamily | SSF47459 | 9.81E-19 | 432 | 498 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.0E-10 | 433 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.1E-18 | 433 | 492 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.8E-16 | 435 | 484 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000084 | anatomy | plant sperm cell | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 590 aa Download sequence Send to blast |
MNIGRLVWNE DDKAIVASLL GKRALDYLLS NSVSNANLLM TLGSDENLQN KLSDLVERPN 60 ASNFSWNYAI FWQISRSKAG DLVLCWGDGY CREPKEGEKS EIVRILSMGR EEETHQTMRK 120 RVLQKLHDLF GGSEEENCAL GLDRVTDTEM FLLSSMYFSF PRGEGGPGKC FASAKPVWLS 180 DVVNSGSDYC VRSFLAKSAG IQTVVLVPTD LGVVELGSTS CLPESEDSIL SIRSLFTSSL 240 PPVRAVALPV TVAEKIDDNR TKIFGKDLHN SGFLQHHQHH QQQQQQPPQQ QQHRQFREKL 300 TVRKMDDRAP KRLDAYPNNG NRFMFSNPGT NNNTLLSPTW VQPENYTRPI NVKEVPSTDE 360 FKFLPLQQSS QRLLPPAQMQ IDFSAASSRA SENNSDGEGG GEWADAVGAD ESGNNRPRKR 420 GRRPANGRAE ALNHVEAERQ RREKLNQRFY ALRSVVPNIS KMDKASLLGD AVSYINELHA 480 KLKVMEAERE RLGYSSNPPI SLDSDINVQT SGEDVTVRIN CPLESHPASR IFHAFEESKV 540 EVINSNLEVS QDTVLHTFVV KSEELTKEKL ISALSREQTN SVQSRTSSGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 4e-26 | 427 | 489 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 261050_at | 0.0 | ||||
| Expression Atlas | AT1G01260 | - | ||||
| AtGenExpress | AT1G01260 | - | ||||
| ATTED-II | AT1G01260 | - | ||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | 26531826 | Download |
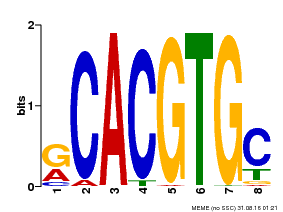
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G01260.3 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT1G51600 | |||||
| IntAct | Search Q9LNJ5 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G01260 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC023628 | 0.0 | AC023628.3 Arabidopsis thaliana chromosome I BAC F6F3 genomic sequence, complete sequence. | |||
| GenBank | AF251698 | 0.0 | AF251698.1 Arabidopsis thaliana putative transcription factor BHLH13 mRNA, complete cds. | |||
| GenBank | AY079012 | 0.0 | AY079012.1 Arabidopsis thaliana At1g01260/F6F3_25 mRNA, complete cds. | |||
| GenBank | AY120752 | 0.0 | AY120752.1 Arabidopsis thaliana transcription factor MYC7E, putative (At1g01260) mRNA, complete cds. | |||
| GenBank | BT004517 | 0.0 | BT004517.1 Arabidopsis thaliana At1g01260/F6F3_25 gene, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001077440.1 | 0.0 | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | ||||
| Refseq | NP_001184883.1 | 0.0 | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | ||||
| Refseq | NP_171634.1 | 0.0 | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | ||||
| Swissprot | Q9LNJ5 | 0.0 | BH013_ARATH; Transcription factor bHLH13 | ||||
| TrEMBL | A0A178WJW5 | 0.0 | A0A178WJW5_ARATH; JAM2 | ||||
| STRING | AT1G01260.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G01260.3 |
| Entrez Gene | 839545 |
| iHOP | AT1G01260 |
| wikigenes | AT1G01260 |



