| Signature Domain? help Back to Top |
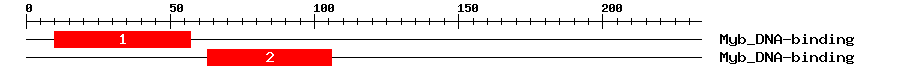 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 56.9 | 4.7e-18 | 10 | 57 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd ll +++ ++G g W++I+ + g++R++k+c++rw++yl
676716888 10 KGAWTAEEDNLLRQCIDKYGEGKWNKIPLRAGLNRCRKSCRLRWLNYL 57
79********************************************97 PP
|
| 2 | Myb_DNA-binding | 41.4 | 3.2e-13 | 63 | 106 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g+++ +E +ll++++k+lG++ W++Ia ++ gRt+ ++k++w++
676716888 63 KGKFSVDEVDLLLRLHKLLGNR-WSLIAGRLS-GRTADDVKNFWNT 106
79********************.*********.***********97 PP
|
| Publications
? help Back to Top |
- Zhang Y, et al.
Pathway engineering for phenolic acid accumulations in Salvia miltiorrhiza by combinational genetic manipulation.
Metab. Eng., 2014. 21: p. 71-80
[PMID:24269612] - Schnaubelt D, et al.
Low glutathione regulates gene expression and the redox potentials of the nucleus and cytosol in Arabidopsis thaliana.
Plant Cell Environ., 2015. 38(2): p. 266-79
[PMID:24329757] - Liu W, et al.
Synthetic TAL effectors for targeted enhancement of transgene expression in plants.
Plant Biotechnol. J., 2014. 12(4): p. 436-46
[PMID:24373379] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Ilk N,Ding J,Ihnatowicz A,Koornneef M,Reymond M
Natural variation for anthocyanin accumulation under high-light and low-temperature stress is attributable to the ENHANCER OF AG-4 2 (HUA2) locus in combination with PRODUCTION OF ANTHOCYANIN PIGMENT1 (PAP1) and PAP2.
New Phytol., 2015. 206(1): p. 422-35
[PMID:25425527] - Li T, et al.
Jasmonic acid enhancement of anthocyanin accumulation is dependent on phytochrome A signaling pathway under far-red light in Arabidopsis.
Biochem. Biophys. Res. Commun., 2014. 454(1): p. 78-83
[PMID:25450360] - Shin DH, et al.
Identification of genes that may regulate the expression of the transcription factor production of anthocyanin pigment 1 (PAP1)/MYB75 involved in Arabidopsis anthocyanin biosynthesis.
Plant Cell Rep., 2015. 34(5): p. 805-15
[PMID:25604992] - Tian J, et al.
McMYB10 regulates coloration via activating McF3'H and later structural genes in ever-red leaf crabapple.
Plant Biotechnol. J., 2015. 13(7): p. 948-61
[PMID:25641214] - Wang L,ZengJ HQ,Song J,Feng SJ,Yang ZM
miRNA778 and SUVH6 are involved in phosphate homeostasis in Arabidopsis.
Plant Sci., 2015. 238: p. 273-85
[PMID:26259194] - Lotkowska ME, et al.
The Arabidopsis Transcription Factor MYB112 Promotes Anthocyanin Formation during Salinity and under High Light Stress.
Plant Physiol., 2015. 169(3): p. 1862-80
[PMID:26378103] - Boter M, et al.
FILAMENTOUS FLOWER Is a Direct Target of JAZ3 and Modulates Responses to Jasmonate.
Plant Cell, 2015. 27(11): p. 3160-74
[PMID:26530088] - He X,Li Y,Lawson D,Xie DY
Metabolic engineering of anthocyanins in dark tobacco varieties.
Physiol Plant, 2017. 159(1): p. 2-12
[PMID:27229540] - Broeckling BE,Watson RA,Steinwand B,Bush DR
Intronic Sequence Regulates Sugar-Dependent Expression of Arabidopsis thaliana Production of Anthocyanin Pigment-1/MYB75.
PLoS ONE, 2016. 11(6): p. e0156673
[PMID:27248141] - Li Y, et al.
Two IIIf Clade-bHLHs from Freesia hybrida Play Divergent Roles in Flavonoid Biosynthesis and Trichome Formation when Ectopically Expressed in Arabidopsis.
Sci Rep, 2016. 6: p. 30514
[PMID:27465838] - Lee WJ, et al.
Drastic anthocyanin increase in response to PAP1 overexpression in fls1 knockout mutant confers enhanced osmotic stress tolerance in Arabidopsis thaliana.
Plant Cell Rep., 2016. 35(11): p. 2369-2379
[PMID:27562381] - Khare D, et al.
Root avoidance of toxic metals requires the GeBP-LIKE 4 transcription factor in Arabidopsis thaliana.
New Phytol., 2017. 213(3): p. 1257-1273
[PMID:27768815] - Li S, et al.
MYB75 Phosphorylation by MPK4 Is Required for Light-Induced Anthocyanin Accumulation in Arabidopsis.
Plant Cell, 2016. 28(11): p. 2866-2883
[PMID:27811015] - Sawano H, et al.
Double-stranded RNA-binding protein DRB3 negatively regulates anthocyanin biosynthesis by modulating PAP1 expression in Arabidopsis thaliana.
J. Plant Res., 2017. 130(1): p. 45-55
[PMID:27995376] - Zou B, et al.
Calmodulin-binding protein CBP60g functions as a negative regulator in Arabidopsis anthocyanin accumulation.
PLoS ONE, 2017. 12(3): p. e0173129
[PMID:28253311] - Zhang Y, et al.
GA-DELLA pathway is involved in regulation of nitrogen deficiency-induced anthocyanin accumulation.
Plant Cell Rep., 2017. 36(4): p. 557-569
[PMID:28275852] - Mondal SK,Roy S
Genome-wide sequential, evolutionary, organizational and expression analyses of phenylpropanoid biosynthesis associated MYB domain transcription factors in Arabidopsis.
J. Biomol. Struct. Dyn., 2018. 36(6): p. 1577-1601
[PMID:28490275] - Xu Z,Mahmood K,Rothstein SJ
ROS Induces Anthocyanin Production Via Late Biosynthetic Genes and Anthocyanin Deficiency Confers the Hypersensitivity to ROS-Generating Stresses in Arabidopsis.
Plant Cell Physiol., 2017. 58(8): p. 1364-1377
[PMID:28586465] - Park JJ,Dempewolf E,Zhang W,Wang ZY
RNA-guided transcriptional activation via CRISPR/dCas9 mimics overexpression phenotypes in Arabidopsis.
PLoS ONE, 2017. 12(6): p. e0179410
[PMID:28622347] - Lowder LG,Paul JW,Qi Y
Multiplexed Transcriptional Activation or Repression in Plants Using CRISPR-dCas9-Based Systems.
Methods Mol. Biol., 2017. 1629: p. 167-184
[PMID:28623586] - Kataya ARA, et al.
PLATINUM SENSITIVE 2 LIKE impacts growth, root morphology, seed set, and stress responses.
PLoS ONE, 2017. 12(7): p. e0180478
[PMID:28678890] - Duan S, et al.
Functional characterization of a heterologously expressed Brassica napus WRKY41-1 transcription factor in regulating anthocyanin biosynthesis in Arabidopsis thaliana.
Plant Sci., 2018. 268: p. 47-53
[PMID:29362083]
|




