 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.009G088100.1.p | ||||||||
| Common Name | Sb09g008120, SORBIDRAFT_09g008120 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 588aa MW: 64288.5 Da PI: 6.5952 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 43 | 7.9e-14 | 441 | 486 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+iLe Av +I +L
Sobic.009G088100.1.p 441 SHVEAERQRREKLNKRFCALRAIVPNiS------KMDKASILEDAVMHIGDL 486
799***********************66......***************988 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.3E-47 | 54 | 243 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| CDD | cd00083 | 3.21E-14 | 435 | 491 | No hit | No description |
| PROSITE profile | PS50888 | 16.622 | 437 | 486 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 7.72E-18 | 439 | 507 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 3.8E-18 | 441 | 503 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.1E-11 | 441 | 486 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-13 | 443 | 492 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 588 aa Download sequence Send to blast |
MKTEVEEVSI GQFWTEEDKA LCSSVLGSDA FTYLTKCGGA ISEGLVAASV LADLQNKLQN 60 LVEADDQSIR WDYAIFWQLS RTKSGAIVLG WGDGSCREPH DSEIGFATSM SVDDASLVTR 120 QKMRKRVLQR LHTAFAGADE EDYAPGIDQV TNTEIFFLAS MYFAFPRHVG GPGKVFGAEA 180 PLWIPNNKHN VSPANYCYRG FLANAAGFKT IVLVPFKAGV LEVGSMQNVP ESAEALQTIR 240 SMFLGTCSDR IAIEKHEDSS SVQISPGLTK IFGKDLSIRQ PSDSKGADPS KVDESSMIAQ 300 KSGGGESRLL PNLRKGLQNF SWSQSRGLSS DQQKFGNGIL VETSETANCS NGASHSTTVS 360 PFQLQNPQQI LAQQASQPRG PMQIDFRVGS SSKFGVLISQ KAMLDVDKGN ANDLFDEERE 420 GRQPRKRERK PTNGREEPPL SHVEAERQRR EKLNKRFCAL RAIVPNISKM DKASILEDAV 480 MHIGDLKKKL EKLEAERDQL PEQTPGPEVD IQVVQGEILV RAVSQIENHP IQKVLQAFED 540 AEVKVGESKV TANNGTVVHS FVIKSPGSEQ HTRKKLLASI SNATSAV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_B | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_E | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_F | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_G | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_I | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_M | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| 5gnj_N | 2e-22 | 434 | 511 | 2 | 78 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.7805 | 0.0 | root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
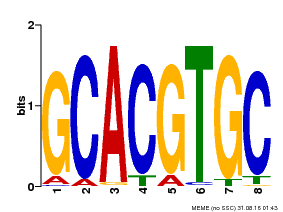
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.009G088100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002440844.1 | 0.0 | transcription factor bHLH13 | ||||
| TrEMBL | C5YV93 | 0.0 | C5YV93_SORBI; Uncharacterized protein | ||||
| STRING | Sb09g008120.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 | Representative plant | OGRP2531 | 11 | 23 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 4e-69 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.009G088100.1.p |
| Entrez Gene | 8066652 |



