 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_01492.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 404aa MW: 45114.9 Da PI: 6.6572 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 39.2 | 1.1e-12 | 80 | 102 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
e C+f+ rtG CkyG++CkF+H+
Rsa1.0_01492.1_g00001.1 80 EDCSFYIRTGSCKYGSSCKFNHP 102
89********************9 PP
| |||||||
| 2 | zf-CCCH | 30 | 8.9e-10 | 130 | 150 | 5 | 25 |
--SGGGGTS--TTTTT-SS-S CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaH 25
+C++++rtG CkyG++C+F+H
Rsa1.0_01492.1_g00001.1 130 ECKYYFRTGGCKYGETCRFSH 150
6******************** PP
| |||||||
| 3 | zf-CCCH | 37.7 | 3.5e-12 | 173 | 197 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
++++C+f++r+G Ck+G+ CkF+H+
Rsa1.0_01492.1_g00001.1 173 GEKECPFYMRNGSCKFGGDCKFNHP 197
7899********************9 PP
| |||||||
| 4 | zf-CCCH | 25.2 | 2.8e-08 | 305 | 330 | 2 | 27 |
-S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHgp 27
++++C+++ +tG Ck+ ++Ck++H++
Rsa1.0_01492.1_g00001.1 305 DQPECSYYVKTGDCKFKSKCKYHHPK 330
689*********************96 PP
| |||||||
| 5 | zf-CCCH | 33 | 9.9e-11 | 351 | 375 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
+++ C +++r+G+Ck+G+ C+F H+
Rsa1.0_01492.1_g00001.1 351 DQSMCTHYSRYGICKFGPACRFDHS 375
5678********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF90229 | 1.83E-6 | 74 | 104 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.122 | 76 | 104 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 3.0E-5 | 76 | 103 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.7E-9 | 80 | 102 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.8E-5 | 82 | 101 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.549 | 125 | 153 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.0E-4 | 125 | 152 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 2.09E-6 | 127 | 154 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.8E-7 | 130 | 150 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.5E-5 | 131 | 152 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.601 | 171 | 199 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 2.8E-6 | 171 | 198 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 8.5E-10 | 173 | 197 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 5.76E-6 | 173 | 199 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 7.8E-4 | 177 | 196 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.751 | 303 | 331 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.0092 | 303 | 330 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.2E-5 | 305 | 330 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 2.88E-5 | 307 | 333 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.906 | 349 | 377 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.019 | 349 | 376 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 6.41E-6 | 350 | 377 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 8.4E-5 | 353 | 375 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.4E-7 | 355 | 375 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 404 aa Download sequence Send to blast |
MSKPDPDPKQ PSGSSSEKDP VAADHITEQE LSEDLRNVVL LDEATEELNV PISVSDGDQQ 60 EEEEGREKRM MVYPVRPDAE DCSFYIRTGS CKYGSSCKFN HPLRRNPQIG REKVRERERD 120 EDVENPRLME CKYYFRTGGC KYGETCRFSH TKQQTSLPSR PELNFLGLPI RPGEKECPFY 180 MRNGSCKFGG DCKFNHPDPK AVGGGVDSPL FRGNNGAPKD APQASSTSWS SSRHMNGTGT 240 GGTAPFIPLM YSQNRGVSPH TPEWSGYQAP SAYPPERSVL PPSTYSAETS SFSQLPAEEF 300 PERPDQPECS YYVKTGDCKF KSKCKYHHPK NRLPKQSPSS FNDKGLPLRP DQSMCTHYSR 360 YGICKFGPAC RFDHSIPPTF SSSSSQTVDT PQLGGNANEN DSWN |
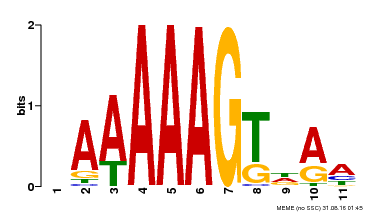
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_01492.1_g00001.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228790 | 1e-107 | AK228790.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL16-10-O16. | |||
| GenBank | BT020223 | 1e-107 | BT020223.1 Arabidopsis thaliana At5g63260 mRNA, complete cds. | |||
| GenBank | BT020443 | 1e-107 | BT020443.1 Arabidopsis thaliana At5g63260 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018432864.1 | 0.0 | PREDICTED: zinc finger CCCH domain-containing protein 67 | ||||
| Swissprot | Q5RJC5 | 0.0 | C3H67_ARATH; Zinc finger CCCH domain-containing protein 67 | ||||
| TrEMBL | M4FC03 | 0.0 | M4FC03_BRARP; Uncharacterized protein | ||||
| STRING | Bra038619.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4176 | 27 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.1 | 0.0 | C3H family protein | ||||



