 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 29970.m000987 | ||||||||
| Common Name | LOC8285592, RCOM_1153840 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 339aa MW: 36448.6 Da PI: 7.7705 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 61.5 | 1.7e-19 | 12 | 57 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEd+ l ++v+++G+++W++I++ ++ gR++k+c++rw +
29970.m000987 12 KGPWSPEEDDALQKLVQKHGPRNWSLISKSIP-GRSGKSCRLRWCNQ 57
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 57.1 | 4e-18 | 66 | 108 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++T+eEde +++a++++G++ W+tIar ++ gRt++ +k++w++
29970.m000987 66 AFTPEEDETIIRAHARFGNK-WATIARLLN-GRTDNAIKNHWNST 108
79******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 26.566 | 7 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.28E-32 | 9 | 105 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.6E-17 | 11 | 60 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.9E-19 | 12 | 57 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-26 | 13 | 65 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.53E-16 | 14 | 56 | No hit | No description |
| SMART | SM00717 | 3.8E-15 | 63 | 111 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 21.467 | 64 | 113 | IPR017930 | Myb domain |
| CDD | cd00167 | 2.04E-12 | 66 | 109 | No hit | No description |
| Pfam | PF00249 | 1.3E-15 | 66 | 108 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-23 | 66 | 112 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MAVTGKEVDR IKGPWSPEED DALQKLVQKH GPRNWSLISK SIPGRSGKSC RLRWCNQLSP 60 QVEHRAFTPE EDETIIRAHA RFGNKWATIA RLLNGRTDNA IKNHWNSTLK RKCCSLDEGY 120 DGNLCCSNNN NNNNDSLINT QPLKRSVSAG SGVPVSTGLY MNPGSPSGSD VSDSSYVPVF 180 TSSPHVFRPV ARTGGVVSFV EATSSSNDNN NNNNISDDPP TSLSLSLPGV DSSEVSNRVA 240 ESTPVRVPDS TTISLMPVMN QVPSPATAAA VQQGGSANSG GFMGFTTEFM AVMQEMIRKE 300 VRNYMMEQSG GGGMCFQAAG GDGFRNVVGM NRVGVSKIE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 8e-43 | 11 | 112 | 3 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 7e-43 | 11 | 112 | 3 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-43 | 11 | 112 | 3 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that functions in salt stress response. Acts as negative regulator of NHX7/SOS1 and CBL4/SOS3 induction in response to salt stress (PubMed:23809151). In response to auxin, activates the transcription of the auxin-responsive gene IAA19. The IAA19 transcription activation by MYB73 is enhanced by direct interaction between MYB73 and PYL8 (PubMed:24894996). {ECO:0000269|PubMed:23809151, ECO:0000269|PubMed:24894996}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00476 | DAP | Transfer from AT4G37260 | Download |
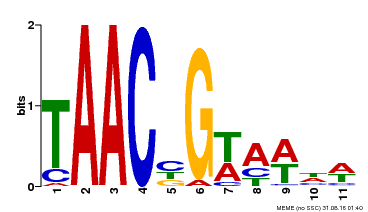
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 29970.m000987 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt stress. {ECO:0000269|PubMed:23809151}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002524926.1 | 0.0 | transcription factor MYB44 | ||||
| Swissprot | O23160 | 1e-105 | MYB73_ARATH; Transcription factor MYB73 | ||||
| TrEMBL | B9SG07 | 0.0 | B9SG07_RICCO; R2r3-myb transcription factor, putative | ||||
| STRING | XP_002524926.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF843 | 34 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G67300.1 | 1e-87 | myb domain protein r1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 29970.m000987 |
| Entrez Gene | 8285592 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



