 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2NG311300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 458aa MW: 46102.8 Da PI: 6.8505 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.9 | 6.4e-12 | 249 | 294 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P+a K +Ka++L + ++Y+k Lq
Pavir.2NG311300.1.p 249 HSIAERLRRERIAERMKALQELVPNA-----NKTDKASMLDEIIDYVKFLQ 294
99***********************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.30E-13 | 242 | 298 | No hit | No description |
| SuperFamily | SSF47459 | 5.5E-17 | 242 | 304 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.39 | 244 | 293 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.9E-16 | 246 | 303 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.1E-9 | 249 | 294 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.5E-13 | 250 | 299 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 458 aa Download sequence Send to blast |
MQPTTREMQA MAAAGQISLD DLRAAGGVHD DFLDQMLGGL QPSAWPELAS AAGAKAPEGG 60 VEAEGMHQQQ QQQFGGGLYD ESALLASRLR QHQISGGGAE SAAAAKQMVL QQLADLRHGH 120 HMLLQGMGRS TVGGGGGDGG MLLPLSLGSG GSGGDVQALL KAAANSAGGE AATVFGGAFA 180 GSLQQQQQQH FQPHPQQQTA PLPGQGFGRG GGASGGASQP QAGASGGTAA AAPPRQRVRA 240 RRGQATDPHS IAERLRRERI AERMKALQEL VPNANKTDKA SMLDEIIDYV KFLQLQVKVL 300 SMSRLGGAAA VAPLVADMSS EGRGGAPPAA AGSDGLAATE QQVAKLMEED MGTAMQYLQG 360 KGLCLMPVSL ASAISSATCH MRPPGLGVAA AAHHMAAMRL PPGMNGAGAD PAAVPASPSM 420 SVLTAQSAMA NGAGGADGEG SHSQQQQPKD AASVSKP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 253 | 258 | RLRRER |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.12702 | 1e-156 | stem | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
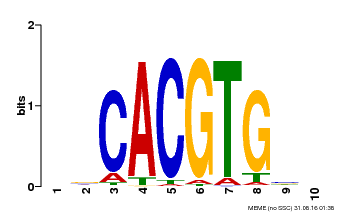
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2NG311300.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold, UV, ethylene (ACC), flagellin, jasmonic acid (JA), and salicylic acid (SA) treatments. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT062076 | 0.0 | BT062076.1 Zea mays full-length cDNA clone ZM_BFb0212P11 mRNA, complete cds. | |||
| GenBank | KJ727938 | 0.0 | KJ727938.1 Zea mays clone pUT6033 bHLH transcription factor (bHLH88) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025799597.1 | 0.0 | glycine, alanine and asparagine-rich protein-like | ||||
| Swissprot | Q9ZUG9 | 8e-79 | BH066_ARATH; Transcription factor bHLH66 | ||||
| TrEMBL | A0A2T7ESB1 | 0.0 | A0A2T7ESB1_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Ba01914.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1419 | 37 | 118 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G24260.1 | 1e-48 | LJRHL1-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2NG311300.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



