 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.8G270000.1.p | ||||||||
| Common Name | PRUPE_ppa004560mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 532aa MW: 59203.8 Da PI: 4.8148 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.9 | 1.8e-16 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+grWT+eEd++l+++++ G g+W++ ++ g+ R++k+c++rw +yl
Prupe.8G270000.1.p 14 KGRWTAEEDQILINYIQTNGEGSWRSLPKNAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.9 | 1.5e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ +++E+++++++++ lG++ W++Ia+ ++ gRt++++k++w+++l
Prupe.8G270000.1.p 67 RGNISAQEEDIIIKLHASLGNR-WSLIASQLP-GRTDNEIKNYWNSHL 112
7899******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-21 | 6 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.678 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.94E-28 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.8E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.2E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.75E-9 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 24.903 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-24 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.2E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.19E-10 | 71 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009813 | Biological Process | flavonoid biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 532 aa Download sequence Send to blast |
MGRASCCNKI GLKKGRWTAE EDQILINYIQ TNGEGSWRSL PKNAGLLRCG KSCRLRWINY 60 LRADLKRGNI SAQEEDIIIK LHASLGNRWS LIASQLPGRT DNEIKNYWNS HLSRKIDTFR 120 RPTTTTSDQM SSLPAAASNN IPSKRRGGRT SRWAMKKSKT YTTTHSTNYT QRHNKRQKDI 180 TNIAADDDEA IALETKTPLP GPNDNIDTMH HDYMVLMTDP VADDHDHHPH HMDDCRVDNL 240 VNHDHDHQRQ EEAGGLAMPA VLMSSTTTIT EEEEEEVEVE KKETRDHDGL CPVNIDQCQK 300 ESHEMLLGPH PHNDENKDDD LDESGGIEFD GGLLGTFNEL IDNVELLQKD PNSNGVLTLS 360 DHQHDHAMGV THEVDQVETT TTCGHLSSSS NEQVCFSSIM SMTATSSSSA SASAAAGSYY 420 FDMEDGQAAA AANGNDRDHN NHILWDWESV VEAGHELWDH DDDKENMLSW LWEEGTCSSS 480 TTSNTTAAAA AASTIDRHLN WEGDTTTDDT SMMRKAVDPD KQNAMVAWLL S* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 9e-24 | 12 | 118 | 5 | 110 | B-MYB |
| 1h8a_C | 9e-24 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
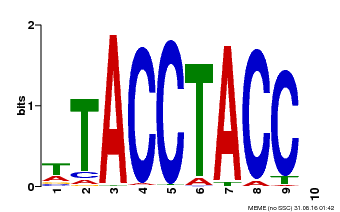
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.8G270000.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU938682 | 0.0 | GU938682.1 Prunus avium R2R3-MYB transcription factor (MYBF1) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020425532.1 | 0.0 | transcription factor MYB29 | ||||
| TrEMBL | A0A251N776 | 0.0 | A0A251N776_PRUPE; Uncharacterized protein | ||||
| STRING | EMJ02170 | 0.0 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47460.1 | 3e-73 | myb domain protein 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.8G270000.1.p |
| Entrez Gene | 18766477 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



