 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00188g00066.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 202aa MW: 23424.2 Da PI: 6.0862 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 47.6 | 3.7e-15 | 87 | 145 | 5 | 63 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
+r rr+++NRe+ArrsR RK+ ++eL v L +eN++L+++l+++++ +++ +e+
Peaxi162Scf00188g00066.1 87 RRKRRMISNRESARRSRMRKQRHLDELWSQVLRLRTENHTLIDKLNQISESHERVLQEN 145
799***********************************************999998886 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 3.1E-16 | 83 | 147 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.266 | 85 | 148 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.0E-11 | 87 | 159 | No hit | No description |
| SuperFamily | SSF57959 | 1.69E-12 | 87 | 137 | No hit | No description |
| Pfam | PF00170 | 6.3E-13 | 87 | 145 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14702 | 5.14E-20 | 88 | 138 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 90 | 105 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 202 aa Download sequence Send to blast |
MIPSEAAATH YFATENPSPL PLDFNFMQNM SSPTFQFSRY LTNLQNYPTS LAVHHDFNNN 60 LPLPSCISSN STSDEADEQQ LRVIDERRKR RMISNRESAR RSRMRKQRHL DELWSQVLRL 120 RTENHTLIDK LNQISESHER VLQENVQLKE EATDLRQMLT DIQLINSPFP DFSDLEEVPC 180 TTAHLKAESS NLSITTSTNL LH |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 99 | 106 | RRSRMRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in somatic embryogenesis. Acts as positive regulator of BHLH109. {ECO:0000269|PubMed:26973252}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
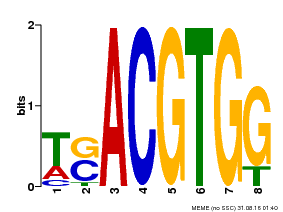
| Motif ID | Method | Source | Motif file |
| MP00647 | PBM | Transfer from LOC_Os02g49560 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016553291.1 | 1e-112 | PREDICTED: basic leucine zipper 43-like | ||||
| Swissprot | Q9FMC2 | 5e-40 | BZP43_ARATH; Basic leucine zipper 43 | ||||
| TrEMBL | A0A1U8F3T3 | 1e-110 | A0A1U8F3T3_CAPAN; basic leucine zipper 43-like | ||||
| TrEMBL | A0A2G2VKG6 | 1e-110 | A0A2G2VKG6_CAPBA; Basic leucine zipper 43 | ||||
| TrEMBL | A0A2G3B6R0 | 1e-110 | A0A2G3B6R0_CAPCH; Basic leucine zipper 43 | ||||
| STRING | XP_009774323.1 | 1e-111 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA947 | 24 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G30530.1 | 3e-47 | basic leucine-zipper 42 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



