 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_120049g0010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 259aa MW: 27784.3 Da PI: 8.5083 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.9 | 4.2e-17 | 18 | 63 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEd l + v++ G+++W++I++ ++ gR++k+c++rw +
MA_120049g0010 18 KGPWSPEEDAALQHFVQKCGPRNWSLISKAIP-GRSGKSCRLRWCNQ 63
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 58.5 | 1.5e-18 | 72 | 114 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++T+eEd +++a++q+G++ W+tIar++ gRt++ +k++w++
MA_120049g0010 72 PFTPEEDATIIRAHAQHGNK-WATIARMLS-GRTDNAIKNHWNST 114
89******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 26.144 | 13 | 68 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.87E-32 | 15 | 111 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-15 | 17 | 66 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.2E-17 | 18 | 63 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-25 | 19 | 71 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.15E-15 | 20 | 62 | No hit | No description |
| SMART | SM00717 | 1.3E-15 | 69 | 117 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 21.278 | 70 | 119 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.2E-15 | 72 | 114 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.44E-13 | 72 | 115 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-24 | 72 | 118 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 259 aa Download sequence Send to blast |
MASMKGKSSG HDEPDRIKGP WSPEEDAALQ HFVQKCGPRN WSLISKAIPG RSGKSCRLRW 60 CNQLSPQVEH RPFTPEEDAT IIRAHAQHGN KWATIARMLS GRTDNAIKNH WNSTLRRRCQ 120 GGGALVIDDE ITSGADGFRK RSLSEDADAS RKLKKLSLGT TATTTTTEPC SSSASDRSDS 180 SLALGSPPVY RPVPRAAPLS LTNEANLNPS HHEASCSSAD PPTSLCLSLP GTDAPAAPTL 240 PQDLPLPSVP CRCLGLLLP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h88_C | 9e-39 | 14 | 119 | 54 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 9e-39 | 14 | 119 | 54 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that functions in salt stress response. Acts as negative regulator of NHX7/SOS1 and CBL4/SOS3 induction in response to salt stress (PubMed:23809151). In response to auxin, activates the transcription of the auxin-responsive gene IAA19. The IAA19 transcription activation by MYB73 is enhanced by direct interaction between MYB73 and PYL8 (PubMed:24894996). {ECO:0000269|PubMed:23809151, ECO:0000269|PubMed:24894996}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00476 | DAP | Transfer from AT4G37260 | Download |
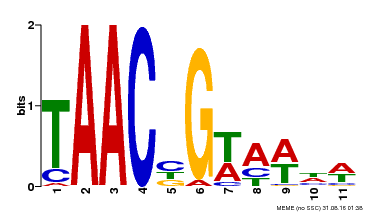
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt stress. {ECO:0000269|PubMed:23809151}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT101188 | 0.0 | BT101188.1 Picea glauca clone GQ0071_N15 mRNA sequence. | |||
| GenBank | EF601069 | 0.0 | EF601069.1 Picea glauca R2R3-MYB transcription factor MYB6 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002285015.1 | 2e-80 | PREDICTED: transcription factor MYB44 | ||||
| Swissprot | O23160 | 4e-68 | MYB73_ARATH; Transcription factor MYB73 | ||||
| TrEMBL | A5JYF0 | 1e-154 | A5JYF0_PICGL; R2R3-MYB transcription factor MYB6 | ||||
| STRING | VIT_18s0001g09850.t01 | 8e-80 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G50060.1 | 3e-65 | myb domain protein 77 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



