 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os03g19120.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 359aa MW: 38768.1 Da PI: 5.0777 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56 | 9.1e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWTteEde+l+ +++++G g+W++ ++ g+ R++k+c++rw +yl
LOC_Os03g19120.1 14 RGRWTTEEDEKLAGYIAKHGEGSWRSLPKNAGLLRCGKSCRLRWINYL 61
8*******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 42.9 | 1.1e-13 | 67 | 139 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT..........................-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg..........................tWktIartmgkgRtlkqcksrwqkyl 48
rg+ + +E++ ++++++ lG++ +W++Ia++++ gRt++++k++w+++l
LOC_Os03g19120.1 67 RGNISNQEEDVIIKLHATLGNRksyvvkrmdyvclgardycfqqnthvRWSLIASHLP-GRTDNEIKNYWNSHL 139
788999****************************************************.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-21 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.437 | 9 | 65 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.7E-11 | 13 | 63 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 8.81E-21 | 14 | 65 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.0E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.59E-10 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-20 | 65 | 143 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.8E-13 | 66 | 141 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 10.192 | 66 | 143 | IPR017930 | Myb domain |
| Pfam | PF00249 | 3.4E-11 | 67 | 139 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.42E-7 | 71 | 139 | No hit | No description |
| SuperFamily | SSF46689 | 8.81E-21 | 111 | 139 | IPR009057 | Homeodomain-like |
| Pfam | PF06640 | 2.5E-82 | 157 | 358 | IPR010588 | Myb-related protein P, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 359 aa Download sequence Send to blast |
MGRAPCCEKV GLKRGRWTTE EDEKLAGYIA KHGEGSWRSL PKNAGLLRCG KSCRLRWINY 60 LRAGVKRGNI SNQEEDVIIK LHATLGNRKS YVVKRMDYVC LGARDYCFQQ NTHVRWSLIA 120 SHLPGRTDNE IKNYWNSHLS RQIHTYRRTY TAPSEAAVTI DVTKLQAAGK RRGGRTAGQS 180 RKGDKKRAED DPPKETAAAD TPLPESSPRR AQSDEARSGS VVVDPEEPSS QPNNGSSGGG 240 GGTPDGPCSE ETATGPTSLD PMEMGLWEVE SEFAEMEALL CGGVAPDGPG IPGLEPLDVA 300 AQADDLLDMD WDGFAADLWG DPAQRGGLVQ DAGEPNGSMG CSSDELESFA SWLLSDSC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-18 | 12 | 145 | 5 | 110 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | Q10MM2 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor postulated to regulate the biosynthetic pathway of a flavonoid-derived pigment in certain floral tissues. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
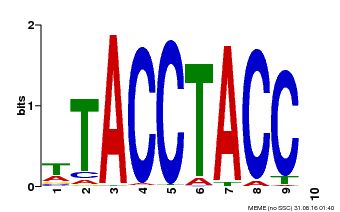
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os03g19120.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os03g19120 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC137267 | 0.0 | AC137267.2 Genomic sequence for Oryza sativa, Nipponbare strain, clone OSJNBa0032G21, from chromosome 3, complete sequence. | |||
| GenBank | AF474141 | 0.0 | AF474141.1 Oryza sativa typical A-type R2R3 Myb protein gene, complete cds. | |||
| GenBank | AP014959 | 0.0 | AP014959.1 Oryza sativa Japonica Group DNA, chromosome 3, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015629329.1 | 0.0 | myb-related protein P | ||||
| Swissprot | P27898 | 2e-86 | MYBP_MAIZE; Myb-related protein P | ||||
| TrEMBL | Q10MM2 | 0.0 | Q10MM2_ORYSJ; P protein C-terminus containing protein | ||||
| STRING | OGLUM03G14490.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47460.1 | 3e-62 | myb domain protein 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os03g19120.1 |



