 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | gw1.6.1363.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 280aa MW: 30331.3 Da PI: 10.5768 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.6 | 3.8e-15 | 29 | 73 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +W++eE+ l+v+ k++G W++I +++g ++t+ q++s+ qk+
gw1.6.1363.1 29 REKWSEEEHALFVESLKKYGRA-WRKIEAHIG-TKTAVQIRSHAQKF 73
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-14 | 23 | 79 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.513 | 24 | 78 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-9 | 25 | 76 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-15 | 27 | 76 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 9.3E-13 | 28 | 76 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-12 | 29 | 72 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.97E-10 | 31 | 74 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 280 aa Download sequence Send to blast |
TETTVEAAAA AQDAAPVKVR KPYTITKKRE KWSEEEHALF VESLKKYGRA WRKIEAHIGT 60 KTAVQIRSHA QKFFSKLQKE QHARAANGSG DGNAQSDGKK RAASASAAGG NKKTKTGLDK 120 RELEIEIPPA RPKRKPAHPY PKKATSQQPS RGSGDRNSGG TGKSFQTEQK MVELGDNAEA 180 IANASSSAAV AAVLSVASDK MQNNLQQDIR ASYFGISPPP MAGMFPQAGL FPMHNMMNPY 240 LGMANPAAGA QNPPQMSNPQ QFVNYINFLT NMWQHNAAVM |
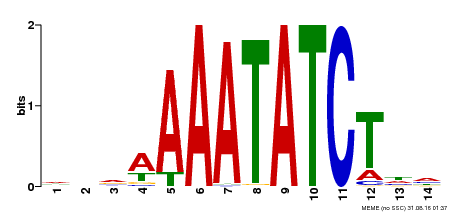
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001418464.1 | 1e-115 | predicted protein, partial | ||||
| TrEMBL | A4RYW3 | 1e-114 | A4RYW3_OSTLU; Uncharacterized protein (Fragment) | ||||
| STRING | ABO96757 | 1e-115 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP454 | 14 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 8e-31 | MYB_related family protein | ||||



