 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G17300.1 | ||||||||
| Common Name | MKP11.15, RVE1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 387aa MW: 42980.9 Da PI: 8.216 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.7 | 2e-16 | 55 | 99 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT eE++++v+a k++G W++I +++g +t+ q++s+ qk+
AT5G17300.1 55 RERWTDEEHKKFVEALKLYGRA-WRRIEEHVG-SKTAVQIRSHAQKF 99
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.57E-16 | 49 | 104 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.908 | 50 | 104 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.3E-16 | 53 | 102 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 8.0E-13 | 54 | 102 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.7E-13 | 55 | 98 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.4E-9 | 55 | 95 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.40E-10 | 57 | 100 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000016 | anatomy | lateral root primordium | ||||
| PO:0000025 | anatomy | root tip | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000039 | anatomy | shoot axis vascular system | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0003011 | anatomy | root vascular system | ||||
| PO:0005660 | anatomy | hydathode | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020003 | anatomy | plant ovule | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0020149 | anatomy | quiescent center | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 387 aa Download sequence Send to blast |
MASSPLTANV QGTNASLRNR DEETADKQIQ FNDQSFGGND YAPKVRKPYT ITKERERWTD 60 EEHKKFVEAL KLYGRAWRRI EEHVGSKTAV QIRSHAQKFF SKVAREATGG DGSSVEPIVI 120 PPPRPKRKPA HPYPRKFGNE ADQTSRSVSP SERDTQSPTS VLSTVGSEAL CSLDSSSPNR 180 SLSPVSSASP PAALTTTANA PEELETLKLE LFPSERLLNR ESSIKEPTKQ SLKLFGKTVL 240 VSDSGMSSSL TTSTYCKSPI QPLPRKLSSS KTLPIIRNSQ EELLSCWIQV PLKQEDVENR 300 CLDSGKAVQN EGSSTGSNTG SVDDTGHTEK TTEPETMLCQ WEFKPSERSA FSELRRTNSE 360 SNSRGFGPYK KRKMVTEEEE HEIHLHL |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.21718 | 0.0 | bud| flower | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 42567911 | 0.0 | ||||
| Genevisible | 250099_at | 0.0 | ||||
| Expression Atlas | AT5G17300 | - | ||||
| AtGenExpress | AT5G17300 | - | ||||
| ATTED-II | AT5G17300 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Myb-like transcription factor that regulates hypocotyl growth by regulating free auxin levels in a time-of-day specific manner. | |||||
| UniProt | Morning-phased transcription factor integrating the circadian clock and auxin pathways. Binds to the evening element (EE) of promoters. Does not act within the central clock, but regulates free auxin levels in a time-of-day specific manner. Positively regulates the expression of YUC8 during the day, but has no effect during the night. Negative regulator of freezing tolerance. {ECO:0000269|PubMed:19805390, ECO:0000269|PubMed:23240770}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | 27203113 | Download |
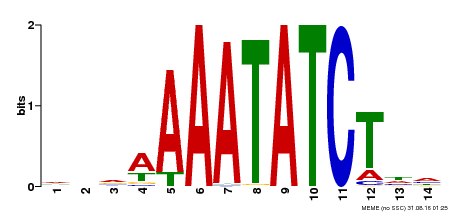
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G17300.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of transcript abundance near subjective dawn. Down-regulated and strongly decreased amplitude of circadian oscillation upon cold treatment. {ECO:0000269|PubMed:19805390, ECO:0000269|PubMed:23240770}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT4G28720(A) | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: No effect on the regulation of core clock associated genes, but shorter hypocotyl length and higher freezing tolerance. Rve1 and rve2 double mutant has no alteration in the period or phase of the clock. Rve1, rve2 and rve7 triple mutant has no alteration in the period or phase of the clock. {ECO:0000269|PubMed:19805390, ECO:0000269|PubMed:23240770}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G17300 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF378860 | 0.0 | AF378860.1 Arabidopsis thaliana AT5g17300/MKP11_15 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_568344.2 | 0.0 | Homeodomain-like superfamily protein | ||||
| Swissprot | F4KGY6 | 0.0 | RVE1_ARATH; Protein REVEILLE 1 | ||||
| TrEMBL | A0A178UEX6 | 0.0 | A0A178UEX6_ARATH; RVE1 | ||||
| STRING | AT5G17300.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6356 | 26 | 44 | Representative plant | OGRP1255 | 15 | 49 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G17300.1 |
| Entrez Gene | 831595 |
| iHOP | AT5G17300 |
| wikigenes | AT5G17300 |



