 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ONIVA03G01790.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 388aa MW: 42790.7 Da PI: 5.1867 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 176.5 | 7.6e-55 | 39 | 168 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk...kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
+ppGfrFhPtdeelv +yL+kkv+++k++l +vi+++d+y++ePwdL++ + e++ewyfFs +d+ky+tg+r+nrat +g+Wkatg+dk+
ONIVA03G01790.1 39 VPPGFRFHPTDEELVGYYLRKKVASQKIDL-DVIRDIDLYRIEPWDLQEhcgIGYDEQSEWYFFSYKDRKYPTGTRTNRATMAGFWKATGRDKA 131
69****************************.9***************953432233667*********************************** PP
NAM 92 vlskkgelvglkktLvfykgrapkgektdWvmheyrle 129
v++ k++l+g++ktLvfykgrap+g+ktdW+mheyrle
ONIVA03G01790.1 132 VHD-KSRLIGMRKTLVFYKGRAPNGQKTDWIMHEYRLE 168
***.999*****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.31E-56 | 34 | 173 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 54.003 | 39 | 188 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.8E-28 | 40 | 167 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:1990110 | Biological Process | callus formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 388 aa Download sequence Send to blast |
MLFILPHTQA SFLPTAREEN IQTMQAQVGD HMDAMESCVP PGFRFHPTDE ELVGYYLRKK 60 VASQKIDLDV IRDIDLYRIE PWDLQEHCGI GYDEQSEWYF FSYKDRKYPT GTRTNRATMA 120 GFWKATGRDK AVHDKSRLIG MRKTLVFYKG RAPNGQKTDW IMHEYRLETD ENAPPQRTAY 180 PARSMVETWD YSLHERNIMS AAAAAAFADP SAAYAQMRRQ HRSGRFKQEA ELDGAATALL 240 HYSSHLAELP QLESPSAAAA PLQPNPSQLA TAGEDDDCKG DNGGRRAKKA RAAGDKVATT 300 TDWRALDKFV ASQLSPGECG SMEATAEAAA AAVAGVSSPL DHGDDDMAAL LFLNSDERDE 360 VDRWTGLLGS GAGASGVDGD LGICVFDK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 2e-48 | 36 | 167 | 12 | 140 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:25148240}. | |||||
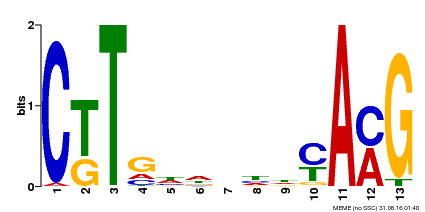
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00266 | DAP | Transfer from AT2G18060 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by chitin (e.g. chitooctaose) (PubMed:17722694). Accumulates in plants exposed to callus induction medium (CIM) (PubMed:17581762). {ECO:0000269|PubMed:17581762, ECO:0000269|PubMed:17722694}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012611 | 0.0 | CP012611.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 3 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015630714.1 | 0.0 | NAC domain-containing protein 37 | ||||
| Refseq | XP_025880076.1 | 0.0 | NAC domain-containing protein 37 | ||||
| Swissprot | Q9SL41 | 1e-115 | NAC37_ARATH; NAC domain-containing protein 37 | ||||
| TrEMBL | A0A0E0GG65 | 0.0 | A0A0E0GG65_ORYNI; Uncharacterized protein | ||||
| STRING | ONIVA03G01790.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3625 | 37 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36160.1 | 1e-107 | NAC domain containing protein 76 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



