 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 32750 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 396aa MW: 43143.6 Da PI: 4.636 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 53.4 | 3.4e-17 | 361 | 392 | 1 | 32 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrk 32
C +Cgt kTp+WR gp+g+ktLCnaCG++y k
32750 361 CLHCGTVKTPQWRMGPEGKKTLCNACGVRYMK 392
99***************************977 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57716 | 8.08E-13 | 354 | 392 | No hit | No description |
| SMART | SM00401 | 4.0E-6 | 355 | 395 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 11.556 | 355 | 390 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 9.9E-14 | 360 | 392 | IPR013088 | Zinc finger, NHR/GATA-type |
| Pfam | PF00320 | 1.1E-14 | 361 | 392 | IPR000679 | Zinc finger, GATA-type |
| CDD | cd00202 | 2.15E-12 | 361 | 390 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 361 | 386 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 396 aa Download sequence Send to blast |
MGGHARARAA PGDFLGAKPS TRDFALEDLF RVDFASGDEG EGEGEKRATI DGGREGDAVR 60 AAATREAREG RAAADDWDAL ARAWDEVEAR APARDYAWEV LNSLPSVGGD IMTCKEDEMD 120 CFAYGGAHSV PYIAPEWLER PNGDVEDGVS DNEDEDEDEG DAHRFTKAMR ESSSFLLDDE 180 DEFMRTVDAP VDFITEPEAF AARREARALA RDAREPAIDF AVKSALLCGD EEEDDFSGQD 240 CSAMADESAY EFQEHFATAA DGRVFRVSSA LVTPSSSERA NVEDSANHRG LTDSEDSDAD 300 LVAHSAKKSP KLIAGSTTKR KRRPDGRGRR AKAYKKIKKD KGVCATDVYV MANGKKMRRG 360 CLHCGTVKTP QWRMGPEGKK TLCNACGVRY MKGIL* |
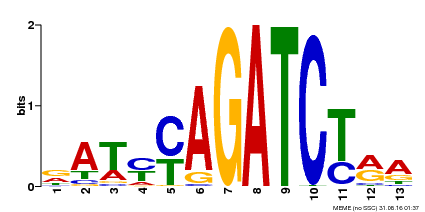
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00530 | DAP | Transfer from AT5G25830 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP000587 | 0.0 | CP000587.1 Ostreococcus lucimarinus CCE9901 chromosome 7, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001418973.1 | 0.0 | predicted protein | ||||
| TrEMBL | A4S0H6 | 0.0 | A4S0H6_OSTLU; Uncharacterized protein | ||||
| STRING | ABO97266 | 0.0 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36240.1 | 5e-15 | GATA transcription factor 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 32750 |



