 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G25830.1 | ||||||||
| Common Name | F18A17.80, GATA12 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 331aa MW: 36489.2 Da PI: 5.2309 | ||||||||
| Description | GATA transcription factor 12 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 55.3 | 9.1e-18 | 221 | 255 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C t kTp+WR gp g+ktLCnaCG++y++ +l
AT5G25830.1 221 CLHCATDKTPQWRTGPMGPKTLCNACGVRYKSGRL 255
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF016992 | 1.0E-79 | 1 | 309 | IPR016679 | Transcription factor, GATA, plant |
| PROSITE profile | PS50114 | 11.635 | 215 | 251 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 8.7E-18 | 215 | 265 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 8.08E-16 | 217 | 279 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.9E-14 | 217 | 252 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 6.54E-15 | 220 | 267 | No hit | No description |
| Pfam | PF00320 | 1.2E-15 | 221 | 255 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 221 | 246 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MEDEAHEFFH TSDFAVDDLL VDFSNDDDEE NDVVADSTTT TTITDSSNFS AADLPSFHGD 60 VQDGTSFSGD LCIPSDDLAD ELEWLSNIVD ESLSPEDVHK LELISGFKSR PDPKSDTGSP 120 ENPNSSSPIF TTDVSVPAKA RSKRSRAAAC NWASRGLLKE TFYDSPFTGE TILSSQQHLS 180 PPTSPPLLMA PLGKKQAVDG GHRRKKDVSS PESGGAEERR CLHCATDKTP QWRTGPMGPK 240 TLCNACGVRY KSGRLVPEYR PAASPTFVLA KHSNSHRKVM ELRRQKEMSR AHHEFIHHHH 300 GTDTAMIFDV SSDGDDYLIH HNVGPDFRQL I |
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 186525744 | 0.0 | ||||
| Genevisible | 246913_at | 0.0 | ||||
| Expression Atlas | AT5G25830 | - | ||||
| AtGenExpress | AT5G25830 | - | ||||
| ATTED-II | AT5G25830 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the vascular cylinder of roots. Expressed in the differentiation zone of the root stele. {ECO:0000269|PubMed:25265867}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a member of the GATA factor family of zinc finger transcription factors. | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. May be involved in the regulation of some light-responsive genes (By similarity). Transcription activator involved in xylem formation. Functions upstream of NAC030/VND7, a master switch of xylem vessel differentiation (PubMed:25265867). {ECO:0000250|UniProtKB:Q8LAU9, ECO:0000269|PubMed:25265867}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00530 | DAP | 27203113 | Download |
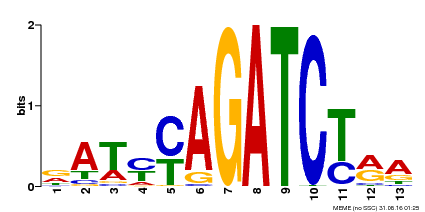
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G25830.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G25830 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB493758 | 0.0 | AB493758.1 Arabidopsis thaliana At5g25830 mRNA for hypothetical protein, partial cds, clone: RAAt5g25830. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_197955.1 | 0.0 | GATA transcription factor 12 | ||||
| Swissprot | P69781 | 0.0 | GAT12_ARATH; GATA transcription factor 12 | ||||
| TrEMBL | A0A178UTT9 | 0.0 | A0A178UTT9_ARATH; GATA transcription factor | ||||
| STRING | AT5G25830.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6625 | 26 | 44 | Representative plant | OGRP68 | 17 | 287 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G25830.1 |
| Entrez Gene | 832652 |
| iHOP | AT5G25830 |
| wikigenes | AT5G25830 |



