 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART03G37400.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 269aa MW: 28625.2 Da PI: 5.1763 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 26.2 | 1.4e-08 | 119 | 163 | 5 | 54 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
h+++Er RR+ri +++ L+el+P+ K + a++L + +Y+k L
OBART03G37400.1 119 HSIAERLRRERIAERMRALQELVPNT-----NKTDRAAMLDEILDYVKFL 163
99***********************7.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 5.23E-15 | 112 | 173 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.00E-10 | 112 | 168 | No hit | No description |
| PROSITE profile | PS50888 | 15.77 | 114 | 163 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.5E-14 | 116 | 174 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 6.0E-6 | 119 | 163 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-9 | 120 | 169 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 269 aa Download sequence Send to blast |
MAGQQPQQQG PPEDDFFDQF FSLTSSFPGA APGGRAAGDQ PFSLALSLDA AAAAEASGSG 60 KRLGVGDDAE GGGSKADRET VQLTGLFPPS KQGGAAVGPQ PPAPRPKVRA RRGQATDPHS 120 IAERLRRERI AERMRALQEL VPNTNKTDRA AMLDEILDYV KFLRLQVKVL SMSRLGGAGA 180 VAQLVADIPL SVKGEASDSG GNQQIWEKWS TDGTERQVAK LMEEDIGAAM QFLQSKALCM 240 MPISLAMAIY DTQQTQDGQP VKHEPNTPS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 123 | 128 | RLRRER |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Required for ovule fertilization. {ECO:0000269|PubMed:15634699}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
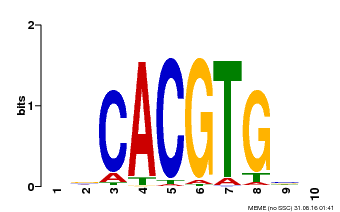
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK068388 | 0.0 | AK068388.1 Oryza sativa Japonica Group cDNA clone:J013145C08, full insert sequence. | |||
| GenBank | AK104412 | 0.0 | AK104412.1 Oryza sativa Japonica Group cDNA clone:006-203-H10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015633087.1 | 1e-158 | transcription factor UNE12 isoform X1 | ||||
| Swissprot | O22768 | 3e-75 | UNE12_ARATH; Transcription factor UNE12 | ||||
| TrEMBL | A0A0D3FQ68 | 0.0 | A0A0D3FQ68_9ORYZ; Uncharacterized protein | ||||
| STRING | OPUNC03G34510.2 | 1e-160 | (Oryza punctata) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G02590.3 | 2e-54 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



