 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR01G36850.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 313aa MW: 33033.1 Da PI: 6.8753 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 124.4 | 1.3e-38 | 80 | 172 | 3 | 102 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssaseceaesssssasnsssgkaaksaak 96
+k+++++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++t++WLlqqa+pai ++tgt++++as+ +++ s ++ +++aa
LPERR01G36850.1 80 PKRSSNKDRHTKVDGRGRRIRMPALCAARIFQLTRELGHKSDGETVQWLLQQAEPAIVAATGTGTIPASALASVAPSL-------PSPNSAAAA 166
6999***************************************************************96662222222.......222222222 PP
TCP 97 skksqk 102
++s++
LPERR01G36850.1 167 LSRSHH 172
222222 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 7.1E-33 | 84 | 297 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 28.039 | 85 | 139 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 313 aa Download sequence Send to blast |
MDPKFPPPPP LNKTEPTTTT TTTTTNQLDH HHQQQPQQQL QIQIHPHQDG GGGKDQHQQQ 60 QEMQVAAAGA GERRVQGLGP KRSSNKDRHT KVDGRGRRIR MPALCAARIF QLTRELGHKS 120 DGETVQWLLQ QAEPAIVAAT GTGTIPASAL ASVAPSLPSP NSAAAALSRS HHHHHHMWPA 180 GFSGASFSAA GGGGGGGDSG GIGGLMQRMG IPGGIDLQGG GGGGHIGFAP MFAGHAAPGL 240 ELGLSQEGHI AGVLAAQSLS QFYHHQVGAG GQLQHHPHQQ HHHQQQHHQQ QQQQQEDGED 300 DRDDGESDEE SGQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the site II motif (3'-TGGGCC/T-5') in the promoter of PCNA-2 and to 3'-GCCCG/A-5' elements in the promoters of cyclin CYCB1-1 and ribosomal protein genes. {ECO:0000269|PubMed:12631321, ECO:0000269|PubMed:16123132}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00636 | PBM | Transfer from Glyma.19G095300 | Download |
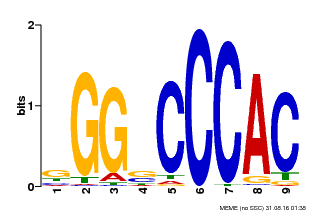
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR01G36850.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072037 | 1e-112 | AK072037.1 Oryza sativa Japonica Group cDNA clone:J013106J07, full insert sequence. | |||
| GenBank | AP004223 | 1e-112 | AP004223.2 Oryza sativa Japonica Group genomic DNA, chromosome 1, BAC clone:B1033B05. | |||
| GenBank | AP004672 | 1e-112 | AP004672.3 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0592G05. | |||
| GenBank | AP014957 | 1e-112 | AP014957.1 Oryza sativa Japonica Group DNA, chromosome 1, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012609 | 1e-112 | CP012609.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 1 sequence. | |||
| GenBank | CT832196 | 1e-112 | CT832196.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCRA111A24, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015621786.1 | 1e-109 | transcription factor TCP20 | ||||
| Swissprot | Q9LSD5 | 9e-47 | TCP20_ARATH; Transcription factor TCP20 | ||||
| TrEMBL | A0A0D9V9Q2 | 0.0 | A0A0D9V9Q2_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR01G36850.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2864 | 32 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27010.1 | 2e-39 | TCP family protein | ||||



