 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.19G095300.1.p | ||||||||
| Common Name | GLYMA_19G095300 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 331aa MW: 35321.1 Da PI: 9.4942 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 116.7 | 2.9e-36 | 66 | 156 | 2 | 89 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssaseceaesssssasnsssg... 88
a+k+++++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti+WLlqqa+p+i+++tgt++++as+ a+ +s s + ss +
Glyma.19G095300.1.p 66 APKRSSNKDRHTKVEGRGRRIRMPALCAARIFQLTRELGHKSDGETIQWLLQQAEPSIIAATGTGTIPASALAAAGNSLSPQGSSLSsal 155
789******************************************************************888444444444433333122 PP
TCP 89 k 89
+
Glyma.19G095300.1.p 156 H 156
0 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 6.6E-32 | 71 | 178 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 28.016 | 72 | 126 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MDPKGSKQQQ PQEVVPKFLS LPQHHYQQQG NSNNNNMGEN KPSEVKDFQI VVAAEKDESK 60 KQQQLAPKRS SNKDRHTKVE GRGRRIRMPA LCAARIFQLT RELGHKSDGE TIQWLLQQAE 120 PSIIAATGTG TIPASALAAA GNSLSPQGSS LSSALHHQHQ KIDELGCGSG GSSRASWQMV 180 GGNLGRPHLG VATTGLWPPH VSGFGFQTAT TTTTTTSSGP SNATLATESS NYLQKIAFPG 240 FDLPTSATNM GHMSFTSILG GAGSQQMPGL ELGLSQDGHI GVLNPQALNQ IYQQMNHQAQ 300 AGRVHQQHQH QQQQHQQTPA KDDSQGSGGQ * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.33531 | 0.0 | cotyledon| epicotyl| hypocotyl| leaf| root| seed coat| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the site II motif (3'-TGGGCC/T-5') in the promoter of PCNA-2 and to 3'-GCCCG/A-5' elements in the promoters of cyclin CYCB1-1 and ribosomal protein genes. {ECO:0000269|PubMed:12631321, ECO:0000269|PubMed:16123132}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
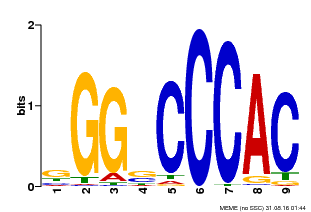
| Motif ID | Method | Source | Motif file |
| MP00636 | PBM | 25215497 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.19G095300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235044 | 0.0 | AC235044.1 Glycine max strain Williams 82 clone GM_WBb0005G15, complete sequence. | |||
| GenBank | AC235265 | 0.0 | AC235265.1 Glycine max strain Williams 82 clone GM_WBb0041K04, complete sequence. | |||
| GenBank | AC236296 | 0.0 | AC236296.1 Glycine max strain Williams 82 clone GM_WBb0094F23, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001304347.2 | 0.0 | transcription factor TCP20-like | ||||
| Refseq | XP_028218369.1 | 0.0 | transcription factor TCP20-like | ||||
| Swissprot | Q9LSD5 | 5e-82 | TCP20_ARATH; Transcription factor TCP20 | ||||
| TrEMBL | A0A445FED1 | 0.0 | A0A445FED1_GLYSO; Transcription factor TCP20 | ||||
| TrEMBL | I1N7U7 | 0.0 | I1N7U7_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA19G26560.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3339 | 32 | 69 | Representative plant | OGRP922 | 14 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27010.1 | 6e-77 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.19G095300.1.p |



