 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_71555.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 317aa MW: 35067.7 Da PI: 4.632 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 61.2 | 1.6e-19 | 65 | 116 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ eq+++Le+ Fe +++++ e+++++A+ lgL rqV vWFqNrRa++k
MLOC_71555.2 65 RRLALEQVRALERSFETDNKLDPERKARIARDLGLHPRQVAVWFQNRRARWK 116
467789*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.9 | 2.1e-41 | 63 | 154 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
kkrrl+ eqv++LE+sFe+++kL+perK+++ar+Lgl+prqvavWFqnrRAR+ktkqlE+d++aL++++dal+++ ++L++++++L++e++e
MLOC_71555.2 63 KKRRLALEQVRALERSFETDNKLDPERKARIARDLGLHPRQVAVWFQNRRARWKTKQLERDFNALRARHDALRADCDALRRDKDALAAEIHE 154
9**************************************************************************************99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00389 | 1.9E-19 | 61 | 122 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.04E-18 | 61 | 120 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 4.30E-17 | 63 | 119 | No hit | No description |
| PROSITE profile | PS50071 | 16.552 | 63 | 118 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 8.9E-17 | 65 | 116 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-19 | 70 | 117 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 4.4E-5 | 89 | 98 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 93 | 116 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 4.4E-5 | 98 | 114 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.9E-16 | 118 | 159 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MKRQIKRPAG SKESPEPGEK LAFAEKENLL AAYMDHEEPE LEEADEEEEE EEERAMSCGL 60 GGKKRRLALE QVRALERSFE TDNKLDPERK ARIARDLGLH PRQVAVWFQN RRARWKTKQL 120 ERDFNALRAR HDALRADCDA LRRDKDALAA EIHELREKLS TKPETAVKAE ATGNVEAAEE 180 RLQQATMVGA AVCKDGSSDS DSSAVFNDEA SPYSGAVFEQ QGFMGFGASF LDTASAAAAT 240 TGCTSLPMLE PKWPSAYPYD ASRSSAYGFT EEWLSGSDAI GNDGSSAFFS EEHVSNLNFG 300 WCSSGAEGFD LQSYCKK |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 110 | 118 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
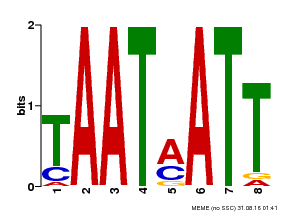
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_71555.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353965 | 0.0 | AK353965.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1004B20. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020190062.1 | 0.0 | homeobox-leucine zipper protein HOX13-like | ||||
| Swissprot | Q10QP3 | 1e-109 | HOX13_ORYSJ; Homeobox-leucine zipper protein HOX13 | ||||
| TrEMBL | F2CQR3 | 0.0 | F2CQR3_HORVV; Predicted protein | ||||
| STRING | MLOC_71555.2 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4283 | 31 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 2e-32 | homeobox 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



