 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.20G155100.1.p | ||||||||
| Common Name | GLYMA_20G155100 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 208aa MW: 22789.3 Da PI: 5.4233 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.4 | 8.5e-19 | 52 | 102 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ y+GVr++ +g+Wv+e+r+p +k+ r++lg+f tae+Aa+a++ a+ +l+g
Glyma.20G155100.1.p 52 PVYRGVRRRD-SGKWVCEVREP---NKKSRIWLGTFPTAEMAARAHDVAAIALRG 102
68*****999.7******9998...347*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.7E-14 | 52 | 102 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-32 | 53 | 112 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.906 | 53 | 110 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.31E-21 | 53 | 112 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.5E-32 | 53 | 116 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 8.04E-30 | 54 | 112 | No hit | No description |
| PRINTS | PR00367 | 3.4E-9 | 54 | 65 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.4E-9 | 76 | 92 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 208 aa Download sequence Send to blast |
MFSINHFSDP DATSPGSGGS SRPALSDEDF FLAASNPKKR AGRKKFRETR HPVYRGVRRR 60 DSGKWVCEVR EPNKKSRIWL GTFPTAEMAA RAHDVAAIAL RGRSACLNFA DSASRLPVPA 120 TAEARDIQKA AAEAAEAFRP GKDDDAVAER VAATATEREE EQEEGMVPEY LRNMVLMSPT 180 HCFGSDEYGS ADVEFDDAEV SLWSYSI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 4e-15 | 51 | 109 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.36065 | 0.0 | hypocotyl | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates abscisic acid- and dehydration-inducible transcription. CBF/DREB1 factors play a key role in freezing tolerance and cold acclimation. {ECO:0000269|PubMed:11798174, ECO:0000269|PubMed:12376631}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00002 | PBM | Transfer from 475045 | Download |
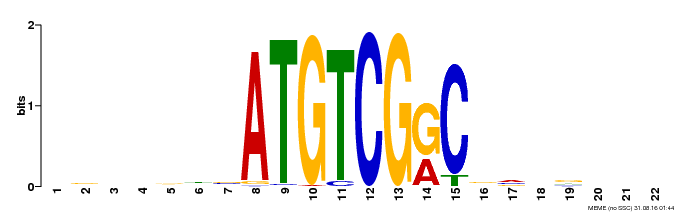
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.20G155100.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high-salt stress, drought stress and abscisic acid (ABA) treatment. {ECO:0000269|PubMed:11798174, ECO:0000269|PubMed:12376631}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF551167 | 0.0 | EF551167.1 Glycine max dehydration-responsive element binding protein 7 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001235037.2 | 1e-151 | dehydration-responsive element binding protein 7 | ||||
| Refseq | XP_028221142.1 | 1e-151 | dehydration-responsive element-binding protein 1E-like | ||||
| Swissprot | Q9FJ93 | 1e-59 | DRE1D_ARATH; Dehydration-responsive element-binding protein 1D | ||||
| TrEMBL | A0A445F5P5 | 1e-150 | A0A445F5P5_GLYSO; Dehydration-responsive element-binding protein 1D | ||||
| TrEMBL | I1NGP6 | 1e-150 | I1NGP6_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA20G29410.1 | 1e-150 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF243 | 33 | 224 | Representative plant | OGRP6 | 16 | 1718 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25480.1 | 3e-56 | dehydration response element B1A | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.20G155100.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



