 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462927065 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 145aa MW: 16291.7 Da PI: 10.8526 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 55.9 | 5.9e-18 | 77 | 111 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C++C ++kTp+WR gp g+ktLCnaCG+++++ +l
462927065 77 CTHCMSSKTPQWRTGPMGPKTLCNACGVRFKSGRL 111
*******************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50114 | 11.451 | 71 | 107 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.8E-17 | 71 | 125 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.85E-15 | 73 | 134 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.3E-14 | 75 | 109 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 4.90E-13 | 76 | 133 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 77 | 102 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 9.3E-16 | 77 | 111 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 145 aa Download sequence Send to blast |
MDVHPPNAAA NSLGDLFPHQ AALVKPENRF TTISEKQWGQ GTNKKQEHGK RRDKKKIKKT 60 PNAIERELPP EPPTRVCTHC MSSKTPQWRT GPMGPKTLCN ACGVRFKSGR LLPEYRPANS 120 PTFVSHIHSN SHKKVLQLRQ GDGHM |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 50 | 58 | RRDKKKIKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00047 | PBM | Transfer from AT1G08000 | Download |
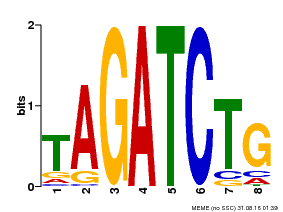
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462927065 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004963249.1 | 2e-51 | GATA transcription factor 2 | ||||
| TrEMBL | A0A0A9VSA1 | 1e-56 | A0A0A9VSA1_ARUDO; Uncharacterized protein | ||||
| STRING | Pavir.Cb00086.1.p | 3e-51 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5962 | 36 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45050.1 | 5e-35 | GATA transcription factor 2 | ||||



