 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.C02592.1.p | ||||||||
| Common Name | EUGRSUZ_C02592, LOC104437555 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 459aa MW: 45954.7 Da PI: 6.7751 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.9 | 6.4e-12 | 255 | 300 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P+a K +Ka++L + ++Y+k Lq
Eucgr.C02592.1.p 255 HSIAERLRRERIAERMKALQELVPNA-----NKTDKASMLDEIIDYVKFLQ 300
99***********************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.32E-13 | 248 | 304 | No hit | No description |
| SuperFamily | SSF47459 | 5.63E-17 | 248 | 310 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.39 | 250 | 299 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.9E-16 | 252 | 309 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.1E-9 | 255 | 300 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.5E-13 | 256 | 305 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 459 aa Download sequence Send to blast |
MQPCSREMQT LSSLLNPQQV SLHDLQGSNG SHPHHGQIQA GGGGAGAGHH HFDPSSSSGD 60 DFLDHMFSGI PSWADHLGTG GDPAAAHHLP MGQNGDVTVD LSEDVGFHGL DEATVLAAKL 120 RQHQIGGGGG GGGGGGAQAA AAAAKAMMLQ QMLMAGGDGS PSSSHFKSAN LGGDSSVQAL 180 FNGFAGSLHG NGQTSNHSQH FSLPQGGSMQ ASPNFSAQAA AAAASGSGGT AGTAAAPGQP 240 RQRVRARRGQ ATDPHSIAER LRRERIAERM KALQELVPNA NKTDKASMLD EIIDYVKFLQ 300 LQVKVLSMSR LGGAAAVAPL VADMSSEGGG GGDGSQANGS GRSSSNGGAS GDSLTVSEHQ 360 VAKLMEEDMG SAMQYLQGKG LCLMPISLAT AISTATCQSR IPLPSPSAHH HHHHHQGPGD 420 GPTSPSVSAL TVQSAALAGG AAAGGGVDAK DAASISKP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 259 | 264 | RLRRER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
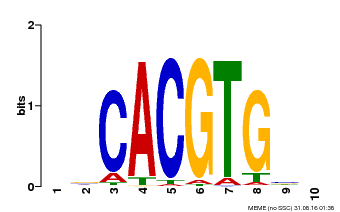
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.C02592.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold, UV, ethylene (ACC), flagellin, jasmonic acid (JA), and salicylic acid (SA) treatments. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP005784 | 2e-32 | AP005784.3 Oryza sativa Japonica Group genomic DNA, chromosome 9, PAC clone:P0014G10. | |||
| GenBank | AP014965 | 2e-32 | AP014965.1 Oryza sativa Japonica Group DNA, chromosome 9, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012617 | 2e-32 | CP012617.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 9 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010048828.1 | 0.0 | PREDICTED: transcription factor bHLH66 | ||||
| Swissprot | Q9ZUG9 | 2e-92 | BH066_ARATH; Transcription factor bHLH66 | ||||
| TrEMBL | A0A059CSL7 | 0.0 | A0A059CSL7_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010048828.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3042 | 27 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G24260.1 | 1e-61 | LJRHL1-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.C02592.1.p |
| Entrez Gene | 104437555 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



