 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC054602.40 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | GRAS | ||||||||
| Protein Properties | Length: 536aa MW: 59059.2 Da PI: 6.1261 | ||||||||
| Description | GRAS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GRAS | 361.8 | 9.9e-111 | 162 | 535 | 2 | 374 |
GRAS 2 velLlecAeavssgdlelaqalLarlselaspdgdpmqRlaayfteALaarlar.svs....elykalppsetseknsseelaalklfsevsPilkf 93
+lL++cAeav+ +d+ +a++lL++l+ +a g+++qR+a++f+++La rl+ + ++ a+ + ++ +e+ al+l ++++P+++f
EcC054602.40 162 LQLLVACAEAVACRDKLQASSLLSELRSRALVFGTSFQRVASCFVQGLADRLSLlQPLgtlgFISPAVVAAASGAAAPDEKEEALRLAYKICPYIQF 258
699************************************************98733221111344444455555555999999************** PP
GRAS 94 shltaNqaIleavegeervHiiDfdisqGl....QWpaLlqaLasRpegpp.slRiTgvgspesgskeeleetgerLakfAeelgvpfefnvlvakr 185
+h++aN Ilea+ege+ vH++D+++s Gl QW L+q+La R ++pp +lRiT++g s+e+ ++g+ L+ +A ++g+++ef+v v+++
EcC054602.40 259 GHFVANTSILEAFEGENFVHVVDLGMSLGLpgghQWRRLIQSLAGRHGQPPrRLRITAIGL----SAEKFCSIGDELSAYALNFGINLEFSV-VETN 350
*************************998644444*************777779********....9*************************9.799* PP
GRAS 186 ledleleeLrvkpgEalaVnlvlqlhrlldesvsleserdevLklvkslsPkvvvvveqeadhnsesFlerflealeyysalfdsleaklpreseer 282
le l+ e++++ +gE+laVn++lqlh +++es + +vL+++ +lsPkv+v+veq+++hn++ Fl rf+eal+yysa+fdsl++ lp+ + r
EcC054602.40 351 LEHLQAEDIKICEGEVLAVNSILQLHCVVKESRGALN---SVLQMIHNLSPKVLVLVEQDSSHNGPFFLGRFMEALHYYSAIFDSLDVMLPKYDTRR 444
*****************************88877777...8******************************************************** PP
GRAS 283 ikvErellgreivnvvacegaerrerhetlekWrerleeaGFkpvplsekaakqaklllrkvk.sdgyrveeesgslvlgWkdrpLvsvSaWr 374
+kvE++++++ei+n+v+ceg +r+erhe++ +Wr+r+++aGF+ +p++ +++qak l k k +gy+v ee+g+lvlgW+++p+v++S+W+
EcC054602.40 445 AKVEQFYFAEEIKNIVSCEGPARVERHERVDQWRRRMSRAGFQLAPIK--MMAQAKQWLGKNKvCEGYTVVEEKGCLVLGWNSKPIVAASCWK 535
**********************************************96..789**********9****************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50985 | 58.683 | 135 | 516 | IPR005202 | Transcription factor GRAS |
| Pfam | PF03514 | 3.4E-108 | 162 | 535 | IPR005202 | Transcription factor GRAS |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006165 | Biological Process | nucleoside diphosphate phosphorylation | ||||
| GO:0006183 | Biological Process | GTP biosynthetic process | ||||
| GO:0006228 | Biological Process | UTP biosynthetic process | ||||
| GO:0006241 | Biological Process | CTP biosynthetic process | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0004550 | Molecular Function | nucleoside diphosphate kinase activity | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0016747 | Molecular Function | transferase activity, transferring acyl groups other than amino-acyl groups | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 536 aa Download sequence Send to blast |
MAPGPYQVPN ELFQNINDDL SHLTIDSVSC LKLASSSSSS SSSLTHYLSP SSIDHEDMLV 60 YCDPEEEEEA RGQKRLKRTY SFAESIGSNS PRAYTGGFGS GGISRSSSMS SLDTFPRLHF 120 RDHIWTYSQR YVAVEAVEEA AAAVSVVGGE EEEGTADGMR LLQLLVACAE AVACRDKLQA 180 SSLLSELRSR ALVFGTSFQR VASCFVQGLA DRLSLLQPLG TLGFISPAVV AAASGAAAPD 240 EKEEALRLAY KICPYIQFGH FVANTSILEA FEGENFVHVV DLGMSLGLPG GHQWRRLIQS 300 LAGRHGQPPR RLRITAIGLS AEKFCSIGDE LSAYALNFGI NLEFSVVETN LEHLQAEDIK 360 ICEGEVLAVN SILQLHCVVK ESRGALNSVL QMIHNLSPKV LVLVEQDSSH NGPFFLGRFM 420 EALHYYSAIF DSLDVMLPKY DTRRAKVEQF YFAEEIKNIV SCEGPARVER HERVDQWRRR 480 MSRAGFQLAP IKMMAQAKQW LGKNKVCEGY TVVEEKGCLV LGWNSKPIVA ASCWKC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3g_A | 1e-57 | 150 | 534 | 12 | 378 | Protein SCARECROW |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
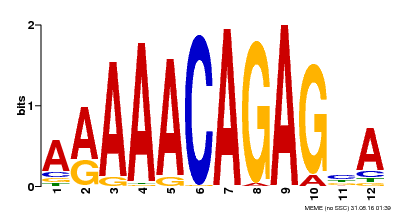
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010039137.1 | 0.0 | PREDICTED: scarecrow-like protein 21 | ||||
| TrEMBL | A0A059A5Q6 | 0.0 | A0A059A5Q6_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010039137.1 | 0.0 | (Eucalyptus grandis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G66350.1 | 6e-71 | RGA-like 1 | ||||



