 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G48430.1 | ||||||||
| Common Name | JMJ12, PKDM9A, REF6, T29H11_50 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1360aa MW: 152629 Da PI: 7.4553 | ||||||||
| Description | relative of early flowering 6 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.9 | 0.00033 | 1243 | 1268 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C+ C++sFs+ +L+ H r+ +
AT3G48430.1 1243 YQCNmeGCTMSFSSEKQLMLHKRNiC 1268
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 15.2 | 6.4e-05 | 1268 | 1290 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H+r+H
AT3G48430.1 1268 CPikGCGKNFFSHKYLVQHQRVH 1290
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 13.6 | 0.00019 | 1326 | 1352 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C dCg++F+ s++ rH r+ H
AT3G48430.1 1326 YVCAepDCGQTFRFVSDFSRHKRKtgH 1352
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 7.6E-18 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.744 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 6.3E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.2E-54 | 200 | 369 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.585 | 203 | 369 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.35E-26 | 216 | 387 | No hit | No description |
| Pfam | PF02373 | 1.2E-37 | 233 | 352 | IPR003347 | JmjC domain |
| SMART | SM00355 | 8 | 1243 | 1265 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.173 | 1266 | 1295 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0081 | 1266 | 1290 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1268 | 1290 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-6 | 1268 | 1294 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.79E-10 | 1282 | 1324 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.6E-9 | 1295 | 1319 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.928 | 1296 | 1325 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 1296 | 1320 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1298 | 1320 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.97E-8 | 1314 | 1348 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-9 | 1320 | 1349 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.406 | 1326 | 1357 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.1 | 1326 | 1352 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1328 | 1352 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0055114 | Biological Process | oxidation-reduction process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0051213 | Molecular Function | dioxygenase activity | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000005 | anatomy | cultured plant cell | ||||
| PO:0000026 | anatomy | primary root tip | ||||
| PO:0000027 | anatomy | lateral root tip | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0003011 | anatomy | root vascular system | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020148 | anatomy | shoot apical meristem | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1360 aa Download sequence Send to blast |
MAVSEQSQDV FPWLKSLPVA PEFRPTLAEF QDPIAYILKI EEEASRYGIC KILPPLPPPS 60 KKTSISNLNR SLAARAAARV RDGGFGACDY DGGPTFATRQ QQIGFCPRKQ RPVQRPVWQS 120 GEEYSFGEFE FKAKNFEKNY LKKCGKKSQL SALEIETLYW RATVDKPFSV EYANDMPGSA 180 FIPLSLAAAR RRESGGEGGT VGETAWNMRA MSRAEGSLLK FMKEEIPGVT SPMVYVAMMF 240 SWFAWHVEDH DLHSLNYLHM GAGKTWYGVP KDAALAFEEV VRVHGYGEEL NPLVTFSTLG 300 EKTTVMSPEV FVKAGIPCCR LVQNPGEFVV TFPGAYHSGF SHGFNFGEAS NIATPEWLRM 360 AKDAAIRRAA INYPPMVSHL QLLYDFVLAL GSRVPTSINP KPRSSRLKDK ARSEGERLTK 420 KLFVQNIIHN NELLSSLGKG SPVALLPQSS SDISVCSDLR IGSHLITNQE NPIQLKCEDL 480 SSDSVVVDLS NGLKDTVSVK EKFTSLCERS RNHLASTEKD TQETLSDAER RKNDAAVALS 540 DQRLFSCVTC GVLSFDCVAI VQPKEAAARY LMSADCSFFN DWTAASGSAN LGQAARSLHP 600 QSKEKHDVNY FYNVPVQTMD HSVKTGDQKT STTSPTIAHK DNDVLGMLAS AYGDSSDSEE 660 EDQKGLVTPS SKGETKTYDQ EGSDGHEEAR DGRTSDFNCQ RLTSEQNGLS KGGKSSLLEI 720 ALPFIPRSDD DSCRLHVFCL EHAAEVEQQL RPFGGINLML LCHPEYPRIE AEAKIVAEEL 780 VINHEWNDTE FRNVTREDEE TIQAALDNVE AKGGNSDWTV KLGVNLSYSA ILSRSPLYSK 840 QMPYNSIIYK AFGRSSPVAS SPSKPKVSGK RSSRQRKYVV GKWCGKVWMS HQVHPFLLEQ 900 DLEGEESERS CHLRVAMDED ATGKRSFPNN VSRDSTTMFG RKYCRKRKIR AKAVPRKKLT 960 SFKREDGVSD DTSEDHSYKQ QWRASGNEEE SYFETGNTAS GDSSNQMSDP HKGIIRHKGY 1020 KEFESDDEVS DRSLGEEYTV RACAASESSM ENGSQHSMYD HDDDDDDIDR QPRGIPRSQQ 1080 TRVFRNPVSY ESEDNGVYQQ SGRISISNRQ ANRMVGEYDS AENSLEERGF CSTGKRQTRS 1140 TAKRIAKTKT VQSSRDTKGR FLQEFASGKK NEELDSYMEG PSTRLRVRHQ KPSRGSLETK 1200 PKKIGKKRSG NASFSRVATE KDVEEKEEEE EEEENEEEEC AAYQCNMEGC TMSFSSEKQL 1260 MLHKRNICPI KGCGKNFFSH KYLVQHQRVH SDDRPLKCPW KGCKMTFKWA WSRTEHIRVH 1320 TGARPYVCAE PDCGQTFRFV SDFSRHKRKT GHSVKKTNKR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 4e-94 | 1223 | 1360 | 3 | 140 | Lysine-specific demethylase REF6 |
| 6a58_A | 4e-94 | 1223 | 1360 | 3 | 140 | Lysine-specific demethylase REF6 |
| 6a59_A | 4e-94 | 1223 | 1360 | 3 | 140 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 945 | 957 | KRKIRAKAVPRKK |
| 2 | 1353 | 1359 | VKKTNKR |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.3169 | 0.0 | flower| root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 79586697 | 0.0 | ||||
| Expression Atlas | AT3G48430 | - | ||||
| AtGenExpress | AT3G48430 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the shoot apical meristem and primary and secondary root tips, and lower expression in cotyledons, leaves and root axis along vascular tissues. Detected in inflorescences, stems and siliques. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18713399}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Relative of Early Flowering 6 (REF6) encodes a Jumonji N/C and zinc finger domain-containing protein that acts as a positive regulator of flowering in an FLC-dependent pathway. REF6 mutants have hyperacetylation of histone H4 at the FLC locus. REF6 interacts with BES1 in a Y2H assay and in vitro. REF6 may play a role in brassinoteroid signaling by affecting histone methylation in the promoters of BR-responsive genes. It is most closely related to the JHDM3 subfamily of JmjN/C proteins. | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
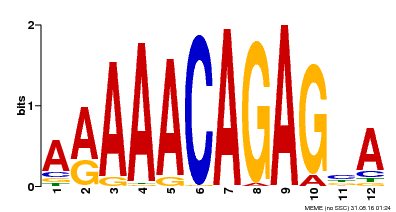
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | SRX1184290 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G48430.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT5G10140(R), AT5G57560(A) | |||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | brassinosteroid | |||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT4G18960, AT1G19350, AT1G24260, AT1G69120 | |||||
| IntAct | Search Q9STM3 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Late-flowering in both long and short days. Brassinosteroid-insensitive phenotype. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G48430 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY664499 | 0.0 | AY664499.1 Arabidopsis thaliana relative of early flowering 6 (REF6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_680116.2 | 0.0 | relative of early flowering 6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A178VKP4 | 0.0 | A0A178VKP4_ARATH; REF6 | ||||
| STRING | AT3G48430.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 | Representative plant | OGRP4401 | 12 | 17 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G48430.1 |
| Entrez Gene | 824002 |
| iHOP | AT3G48430 |
| wikigenes | AT3G48430 |



