 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dusal.0320s00013.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Dunaliellaceae; Dunaliella
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 1081aa MW: 111777 Da PI: 6.9406 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 48.9 | 9.1e-16 | 271 | 299 | 1 | 29 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGly 29
C +Cgt++Tp+WR+gp g+ktLCnaCG++
Dusal.0320s00013.1.p 271 CMQCGTSQTPQWREGPHGPKTLCNACGVR 299
99*************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00439 | 0.0032 | 36 | 161 | IPR001025 | Bromo adjacent homology (BAH) domain |
| PROSITE profile | PS51038 | 12.853 | 36 | 161 | IPR001025 | Bromo adjacent homology (BAH) domain |
| Pfam | PF01426 | 1.1E-8 | 37 | 160 | IPR001025 | Bromo adjacent homology (BAH) domain |
| PROSITE profile | PS50114 | 12.044 | 265 | 298 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 3.5E-11 | 265 | 316 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 4.37E-11 | 268 | 311 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.2E-12 | 268 | 300 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 2.18E-12 | 271 | 320 | No hit | No description |
| Pfam | PF00320 | 1.4E-13 | 271 | 299 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 271 | 296 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1081 aa Download sequence Send to blast |
MQMQVFVWGH PVVSKAEDAG HYPFRQHYQN FWLQGQEYRI GDAVLLESSP EAGSAPYIAQ 60 LLACFRQHSP GGAEEEHFIQ VKWFERATAL NISQHSKEVV ELEETDCNPL GCIGGRCTII 120 RANNWAEASH VAATAVSDPG LEVFFCQGLY NPVTNVLHKY PQLEGGACIG HSKRTMHPTR 180 LPSDEVDSTV VHISSDGYLP GSGSNKRQRL ASPYEPDFLD ASMLDQLLVE EDLRDQQVQH 240 VGHAANIGMC SFEGNLASPP AHGVCVPGRC CMQCGTSQTP QWREGPHGPK TLCNACGVRR 300 VRAQQRSSKS MPSAAARAPK TSHAACMSSA GPQQQQQQQQ QKQPPTSSDK VPATKAPAKA 360 EPKAEAPAST SSRPTRQSAV LAASRNAQFA RTGVFPEIEE RCSSAQVRAA QQQQQQQQQQ 420 SNICRKRRSA SASAAAQQQQ QQQPHQHKAG TVGYQGSGGS SGQGAAARPE AAADSSMWPQ 480 HSCENNSSHL STGSALAALG DGMGTLLSEE GEEAPRDEGR NSSEEPEGGG SLHGGHLHTS 540 TTSATAEASA PQDKPHLRQS SRQQGLAAMA VAARSSHNDL ASFQQQQQQQ QLRGNGVNAA 600 EGSSKASGAS EKQQQQQQGH ADGGGNGVGG SKGRAPAGTA AVAAEPRVPK AGPAPGGASK 660 AATSRAAVGA AAQVGLSSSV AHVPYRVPQV AVFELHERVG SVNGPPAKLE PVVVEGAEAS 720 VAVVGGVTRG KAGGGGSAIS TEGVAKVSHK KGASHKKGAG SAHGGGSKSK AKSKGAKSAA 780 PPAQHENNAT LASQPSATPA LPPAPHTPSP LPGSPPPASP PPPHISPAMA ALPQLPPPSS 840 LSGAKDAAAL MSSGECGVGM GPGFGGEAGE VQDEAMALQL REARSCAEHA AAQVDTADAA 900 VAQAQRVLLE CQEAAERARG DAIAANLRLS ELRHLSGDHQ GGHMAATSLP PPPPHPVGFM 960 PFGPSSFSLL PPLPQQLELL PPAPLLPPPP PPPPPPPPPA SPPHLHPPPP PHLELHAAHQ 1020 PHALLQGPPP PSAGLMPLSG VGLGDDDMGP MCMLPCDGHD VMGHMGDDMG EEMADMMALF 1080 * |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
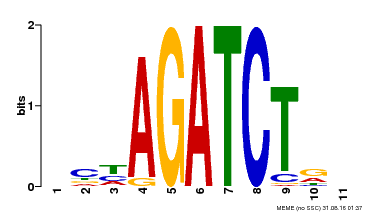
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G26930.1 | 5e-12 | GATA transcription factor 23 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Dusal.0320s00013.1.p |



