 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10020135m | ||||||||
| Common Name | CICLE_v10020130mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 449aa MW: 49689.6 Da PI: 7.2584 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.2 | 1.6e-17 | 26 | 70 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE+++++da k++G g W+ I +++g ++t+ q++s+ qk+
Ciclev10020135m 26 REKWTEEEHQRFLDALKMYGRG-WRQIEEHVG-TKTAVQIRSHAQKF 70
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.17E-16 | 20 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.403 | 21 | 75 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-9 | 23 | 74 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-16 | 24 | 73 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.0E-12 | 25 | 73 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.0E-15 | 26 | 69 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.12E-10 | 28 | 71 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 449 aa Download sequence Send to blast |
MSNSLYSFEN DSLPKVRKPY TITKQREKWT EEEHQRFLDA LKMYGRGWRQ IEEHVGTKTA 60 VQIRSHAQKF FSKVVRESNG SSESSIMPIE IPPPRPKRKP VHPYPRKSVD SLKATSVSNQ 120 QENFTSSNAL VSDKDRQSPT SVLSSFNSDT LGCAASDQQN GCSSPTSCTT EMHSVNLLPI 180 EKENEYVTSI SFPKEEKIST LPAHLSASSN VEELASVSKD SVYPKGDAAA APSCTSIKLF 240 GRTVLVSDSW KPYSLGADSY KSPISKSSQE NLDVDKENFV QSAPSKHLDT HLLLGMVSSN 300 CNPSSHIGPV FQNMELQKKR TNVAEASHNI YLLWDSLYRG APYFHLMPIG ENQAATPLEF 360 SVKDKEILNE RSCSGSSAGS VSELENWEKS SDVAVDSQCP QVCPLSQASP SNCMRGFVPY 420 KRCLAESEIR SSVIVSEERE RQRARVCS* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
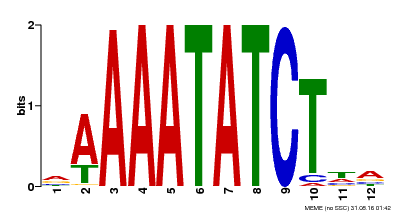
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006441350.1 | 0.0 | protein REVEILLE 7 isoform X1 | ||||
| TrEMBL | V4TXF8 | 0.0 | V4TXF8_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006441350.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 5e-48 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10020135m |
| Entrez Gene | 18050298 |



