 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_40006 | ||||||||
| Common Name | KK1_043067 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 458aa MW: 48431.5 Da PI: 6.4563 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.9 | 6.4e-12 | 242 | 287 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P+a K +Ka++L + ++Y+k Lq
C.cajan_40006 242 HSIAERLRRERIAERMKALQELVPNA-----NKTDKASMLDEIIDYVKFLQ 287
99***********************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.32E-13 | 235 | 291 | No hit | No description |
| SuperFamily | SSF47459 | 5.63E-17 | 235 | 297 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.39 | 237 | 286 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.9E-16 | 239 | 296 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.1E-9 | 242 | 287 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.5E-13 | 243 | 292 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 458 aa Download sequence Send to blast |
MQPCSREMQG LNSLLNSSQI SLQDLHNHNA INQIQNSHMN QHQQQLTPHF DPTSHDDFLD 60 QMLSSCSWPD LAPNKPLWDP NPPTSDETTP SNNDNVAFPY DEHSSLASKF RNHQISTKNN 120 AALILHQQLL MSRAPDSGLL NLPLNLPGND VVDASNFKSP NPGGEASVQA LYNGFTGSLH 180 GAGQASNQTQ HFQHLQGSSN PVHGQNFGGT ANQAPASGAT GGTPAQPRQR VRARRGQATD 240 PHSIAERLRR ERIAERMKAL QELVPNANKT DKASMLDEII DYVKFLQLQV KVLSMSRLGG 300 AAAVAPLVAD MSSEGGGDCI QANGNSNGGG ALAQNSNNNN QTTASTSNDT LTMTEHQVAK 360 LMEEDMGSAM QYLQGKGLCL MPISLATAIS TATCHSRNPL ITAASAGNQI PTNAAHANPA 420 AASNGEGPSS PSMSVLTVQS AVVGNDGAVK DAASVSKP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 246 | 251 | RLRRER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
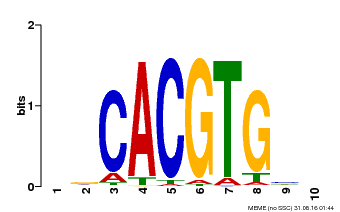
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_40006 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold, UV, ethylene (ACC), flagellin, jasmonic acid (JA), and salicylic acid (SA) treatments. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015035 | 1e-135 | AP015035.1 Vigna angularis var. angularis DNA, chromosome 2, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_029125630.1 | 0.0 | transcription factor bHLH66 | ||||
| Swissprot | Q9ZUG9 | 3e-96 | BH066_ARATH; Transcription factor bHLH66 | ||||
| TrEMBL | A0A151QZU2 | 0.0 | A0A151QZU2_CAJCA; Transcription factor bHLH66 | ||||
| STRING | XP_007138416.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4778 | 33 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G24260.1 | 5e-58 | LJRHL1-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



