 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013628435.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 418aa MW: 46768.7 Da PI: 6.0335 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 39.2 | 1.2e-12 | 89 | 111 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
e C+f+ rtG CkyG++CkF+H+
XP_013628435.1 89 EDCSFYIRTGSCKYGSSCKFNHP 111
89********************9 PP
| |||||||
| 2 | zf-CCCH | 29.9 | 9.3e-10 | 139 | 159 | 5 | 25 |
--SGGGGTS--TTTTT-SS-S CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaH 25
+C++++rtG CkyG++C+F+H
XP_013628435.1 139 ECKYYFRTGGCKYGETCRFSH 159
6******************** PP
| |||||||
| 3 | zf-CCCH | 37.6 | 3.7e-12 | 182 | 206 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
++++C+f++r+G Ck+G+ CkF+H+
XP_013628435.1 182 GEKECPFYMRNGSCKFGGDCKFNHP 206
7899********************9 PP
| |||||||
| 4 | zf-CCCH | 18.4 | 3.7e-06 | 319 | 344 | 2 | 27 |
-S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHgp 27
++++C+++ +tG Ck+ +Ck++H++
XP_013628435.1 319 DQPECSYYVKTGDCKFKYKCKYHHPK 344
689*********************96 PP
| |||||||
| 5 | zf-CCCH | 32.5 | 1.5e-10 | 366 | 389 | 3 | 26 |
S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 3 telCrffartGtCkyGdrCkFaHg 26
++ C +++r+G+Ck+G+ C+F H+
XP_013628435.1 366 QSMCTHYSRYGICKFGPACRFDHS 389
578********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF90229 | 1.96E-6 | 83 | 113 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.821 | 85 | 113 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 5.5E-5 | 85 | 112 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.1E-9 | 89 | 111 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.8E-5 | 91 | 110 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.0E-4 | 134 | 161 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.699 | 134 | 162 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 2.35E-6 | 135 | 162 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 3.0E-7 | 139 | 159 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 3.1E-5 | 140 | 162 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 2.8E-6 | 180 | 207 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.601 | 180 | 208 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.2E-6 | 182 | 207 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 8.9E-10 | 182 | 206 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.076 | 317 | 344 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.577 | 317 | 345 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 8.5E-5 | 321 | 347 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.0072 | 363 | 390 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.662 | 363 | 391 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 4.84E-6 | 363 | 391 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 8.8E-5 | 367 | 389 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 4.4E-7 | 369 | 389 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 418 aa Download sequence Send to blast |
MSKPDPDPKP PSGSSTEKDP VADHVTEELS DDLRNVGLSD EATEELSVPI NVSEGDGETD 60 QKAEDEDEEE GRERVEKRMM VYPVRPDAED CSFYIRTGSC KYGSSCKFNH PVRRKLQIGR 120 EKVKEKEREE NVENPRLMEC KYYFRTGGCK YGETCRFSHT KEHTPLPSRP ELNFLGLPIR 180 PGEKECPFYM RNGSCKFGGD CKFNHPDPTA VGGVDSPLFR GNNGGSFAPK DAPQASSTSW 240 SSPRHMNGTG TAPFIPVMYS QNRGVSPQTP EWSGYQAPSA YPPERNILPP STYSANNSLA 300 ETSSFSQLQS TEEFPERPDQ PECSYYVKTG DCKFKYKCKY HHPKNRLPKQ SPSSFNDKGL 360 PLRPEQSMCT HYSRYGICKF GPACRFDHSI PPTFSSTSSQ TVETPQLGGN GNENDGWN |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
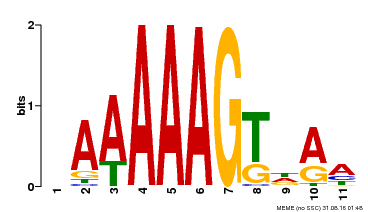
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013628435.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013628435.1 | 0.0 | PREDICTED: zinc finger CCCH domain-containing protein 67 | ||||
| Swissprot | Q5RJC5 | 0.0 | C3H67_ARATH; Zinc finger CCCH domain-containing protein 67 | ||||
| TrEMBL | A0A0D3BFH3 | 0.0 | A0A0D3BFH3_BRAOL; Uncharacterized protein | ||||
| STRING | Bo3g103620.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4176 | 27 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.1 | 0.0 | C3H family protein | ||||



