 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_23919_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 161aa MW: 17802.2 Da PI: 8.8431 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 109.6 | 1.6e-34 | 49 | 103 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+rlrWt+eLHerFv+av+qLGG+++AtPk +l++m+v+gLt++hvkSHLQkYRl
Neem_23919_f_1 49 KQRLRWTHELHERFVDAVAQLGGPDRATPKGVLRVMGVQGLTIYHVKSHLQKYRL 103
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.485 | 46 | 106 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-33 | 48 | 104 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.15E-18 | 48 | 104 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.5E-26 | 49 | 104 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.8E-10 | 51 | 102 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.2E-9 | 135 | 156 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 161 aa Download sequence Send to blast |
MYQPNSVPSS SLVRSNSLVH GQHLDCGSSQ MDPMNGGNNV NNNPSLASKQ RLRWTHELHE 60 RFVDAVAQLG GPDRATPKGV LRVMGVQGLT IYHVKSHLQK YRLAKYLPDS SSDGKKVDKK 120 ETGDMLSSLD GSSGMQITEA LKLQMEVQKR LHEQLEACFC P |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 4e-26 | 49 | 106 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-26 | 49 | 106 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-26 | 49 | 106 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-26 | 49 | 106 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
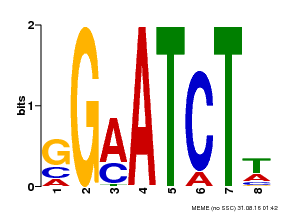
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00628 | PBM | Transfer from PK10342.1 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006449962.1 | 1e-110 | myb family transcription factor PHL7 isoform X1 | ||||
| Refseq | XP_006467243.1 | 1e-110 | myb family transcription factor PHL7 isoform X1 | ||||
| Refseq | XP_024951354.1 | 1e-110 | myb family transcription factor PHL7 isoform X1 | ||||
| Swissprot | Q9SJW0 | 6e-73 | PHL7_ARATH; Myb family transcription factor PHL7 | ||||
| TrEMBL | V4UB20 | 1e-109 | V4UB20_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006467243.1 | 1e-109 | (Citrus sinensis) | ||||
| STRING | XP_006449962.1 | 1e-109 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8753 | 25 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G01060.1 | 2e-53 | G2-like family protein | ||||



