 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aqcoe2G393600.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Ranunculales; Ranunculaceae; Thalictroideae; Aquilegia
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 257aa MW: 29201.1 Da PI: 9.4643 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 52.9 | 5.1e-17 | 201 | 235 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C + kTp+WR gp g+ktLCnaCG++y++ +l
Aqcoe2G393600.2.p 201 CLHCLSEKTPQWRAGPLGPKTLCNACGVRYKSGRL 235
99*****************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57716 | 1.71E-13 | 193 | 247 | No hit | No description |
| SMART | SM00401 | 5.4E-15 | 195 | 249 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 12.044 | 195 | 231 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 2.5E-13 | 198 | 233 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 9.44E-14 | 200 | 250 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 201 | 226 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 8.8E-15 | 201 | 235 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 257 aa Download sequence Send to blast |
MTFSSLPLFL KKKNKNKSPR KLEVFLLSFF LMRMESLESA ALTFETLLDF PSDIDEEPDN 60 NDDDNNIKNP SLSLLTNNQQ INSAELTISH SDAHLHPFPV LEEELEWLNE NSFPDIDTFL 120 SVKPNSNNVA KDQSPVSVLG NSSSSLRHHP VQARSKRRRR RRSSFSDLIG QQWWVSYEPK 180 IKKKNVGATR MSMTSSSGRR CLHCLSEKTP QWRAGPLGPK TLCNACGVRY KSGRLLPEYR 240 PANSPTFSDI RYNDLM* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 153 | 161 | RSKRRRRRR |
| 2 | 155 | 160 | KRRRRR |
| 3 | 155 | 161 | KRRRRRR |
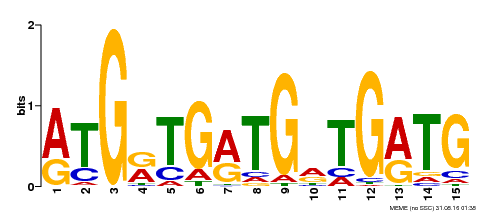
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00377 | DAP | Transfer from AT3G24050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010264014.1 | 5e-50 | PREDICTED: GATA transcription factor 1 | ||||
| TrEMBL | A0A2G5CXZ6 | 1e-158 | A0A2G5CXZ6_AQUCA; Uncharacterized protein | ||||
| STRING | Aquca_034_00184.1 | 1e-159 | (Aquilegia coerulea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 3e-27 | GATA transcription factor 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aqcoe2G393600.2.p |



