 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g006537m | ||||||||
| Common Name | CISIN_1g0021441mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 642aa MW: 70381.6 Da PI: 5.5385 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.7 | 2.6e-29 | 234 | 284 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
orange1.1g006537m 234 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 284
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.484 | 223 | 304 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 2.2E-26 | 237 | 284 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 2.32E-23 | 540 | 629 | No hit | No description |
| Gene3D | G3DSA:3.10.20.240 | 1.3E-27 | 545 | 626 | No hit | No description |
| PROSITE profile | PS51745 | 25.265 | 545 | 627 | IPR000270 | PB1 domain |
| SMART | SM00666 | 9.1E-26 | 545 | 627 | IPR000270 | PB1 domain |
| CDD | cd06407 | 1.30E-39 | 546 | 626 | No hit | No description |
| Pfam | PF00564 | 4.8E-19 | 546 | 626 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 642 aa Download sequence Send to blast |
MGQVCMSTTD VAFYVVDGHM WGFREACVEH HLQKDQGVAG RAFFSLSSCF CKDITQFCKT 60 EYPLVHYARM FGLTSCFAIC LRSTYTGDDD YILEFFLPPA ITDSYEQQTL LGSILATMKQ 120 HFQSLKVASG IDLEDDEGTI EIIEGEATAD KKLNLRMESI RIPQSVRSPP QPHALPNGGE 180 LGQLDIPEQQ LMENFDYMNS RGNAVNVGGN DNPVSLLENK NTRKLSERKR GKTEKSISLE 240 VLQQYFAGSL KDAAKSLGVC PTTMKRICRQ HGISRWPSRK INKVNRSLTK LKRVIESVQG 300 TNGTFGLTSL TTSPLPVAVN SISWPSGLNG SNQQNSPNSK PELLGEKILS PIYKTPGSDG 360 HTELEDRLSG GRMSTHEEHI HEQNALSPEI GKGKNSPKTG SGSREESDGS PTSHGSCQGN 420 PANESAPAKD VLVSSIHEPR FKVGGSLELV FQPVKEMNLS AAFSIPDALV TTEPQEPFGG 480 LLVEDAGSSK DLRNLCPAVA DAIVDERLPE NSCANLPCAE LSPKQHLATL SQTMPRVYSR 540 QEMKSVTIKA TYREDIIRFR ISLSCGILEL KEEVAKRLKL ELGTFDIKYL DDDQEWVLIA 600 CDADLQECLD ISRSSGSNMI RLSIHDIMAN LGSSCESTGE L* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
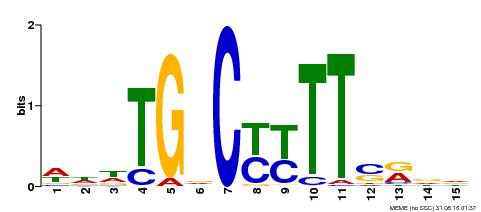
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KX056270 | 0.0 | KX056270.1 Citrus trifoliata transcription factor NLP7 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006439290.2 | 0.0 | LOW QUALITY PROTEIN: protein NLP7 | ||||
| Refseq | XP_006476342.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A067GMH3 | 0.0 | A0A067GMH3_CITSI; Uncharacterized protein | ||||
| STRING | XP_006476342.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006439290.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g006537m |



