 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g005290m | ||||||||
| Common Name | CISIN_1g0021441mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 705aa MW: 77219.6 Da PI: 5.8676 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.6 | 2.9e-29 | 297 | 347 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
orange1.1g005290m 297 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 347
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.484 | 286 | 367 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 2.5E-26 | 300 | 347 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 2.65E-23 | 603 | 692 | No hit | No description |
| SMART | SM00666 | 9.1E-26 | 608 | 690 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.5E-27 | 608 | 689 | No hit | No description |
| PROSITE profile | PS51745 | 25.265 | 608 | 690 | IPR000270 | PB1 domain |
| CDD | cd06407 | 2.97E-39 | 609 | 689 | No hit | No description |
| Pfam | PF00564 | 5.5E-19 | 609 | 689 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 705 aa Download sequence Send to blast |
MLLNQICNEG RQNALAEILE ILSVVCETHK LPLAQTWVPC RHRSVLAYGG GLKKSCSSID 60 GSCMGQVCMS TTDVAFYVVD GHMWGFREAC VEHHLQKDQG VAGRAFFSLS SCFCKDITQF 120 CKTEYPLVHY ARMFGLTSCF AICLRSTYTG DDDYILEFFL PPAITDSYEQ QTLLGSILAT 180 MKQHFQSLKV ASGIDLEDDE GTIEIIEGEA TADKKLNLRM ESIRIPQSVR SPPQPHALPN 240 GGELGQLDIP EQQLMENFDY MNSRGNAVNV GGNDNPVSLL ENKNTRKLSE RKRGKTEKSI 300 SLEVLQQYFA GSLKDAAKSL GVCPTTMKRI CRQHGISRWP SRKINKVNRS LTKLKRVIES 360 VQGTNGTFGL TSLTTSPLPV AVNSISWPSG LNGSNQQNSP NSKPELLGEK ILSPIYKTPG 420 SDGHTELEDR LSGGRMSTHE EHIHEQNALS PEIGKGKNSP KTGSGSREES DGSPTSHGSC 480 QGNPANESAP AKDVLVSSIH EPRFKVGGSL ELVFQPVKEM NLSAAFSIPD ALVTTEPQEP 540 FGGLLVEDAG SSKDLRNLCP AVADAIVDER LPENSCANLP CAELSPKQHL ATLSQTMPRV 600 YSRQEMKSVT IKATYREDII RFRISLSCGI LELKEEVAKR LKLELGTFDI KYLDDDQEWV 660 LIACDADLQE CLDISRSSGS NMIRLSIHDI MANLGSSCES TGEL* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
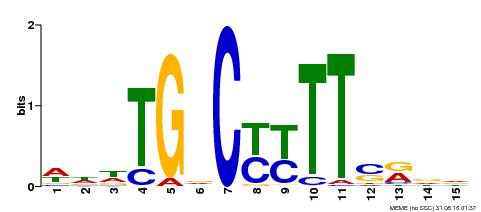
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KX056270 | 0.0 | KX056270.1 Citrus trifoliata transcription factor NLP7 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006476342.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A067GMR4 | 0.0 | A0A067GMR4_CITSI; Uncharacterized protein | ||||
| STRING | XP_006476342.1 | 0.0 | (Citrus sinensis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g005290m |



