 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.136780.1 | ||||||||
| Common Name | Csa_6G012810, LOC101215431 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 411aa MW: 45612.8 Da PI: 4.7961 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.1 | 1.1e-18 | 69 | 119 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
+ y+GVr++ +g+WvAeIr+p + r r +lg+f ta eAa a+++a++ ++g
Cucsa.136780.1 69 CNYRGVRQRT-WGKWVAEIREP---N-RgSRLWLGTFPTAIEAALAYDEAARTMYG 119
56*****999.**********8...3.35***********************9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 3.5E-13 | 69 | 119 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.01E-30 | 69 | 128 | No hit | No description |
| SMART | SM00380 | 1.1E-36 | 70 | 133 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 9.4E-31 | 70 | 128 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.45E-20 | 70 | 127 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 21.509 | 70 | 127 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-9 | 71 | 82 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-9 | 93 | 109 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 411 aa Download sequence Send to blast |
MISETIERKR KSRSRRDRST VAETLAKWKA YNECFDSSNN GGKLIRKAPA KGSKKGCMKG 60 KGGPLNSHCN YRGVRQRTWG KWVAEIREPN RGSRLWLGTF PTAIEAALAY DEAARTMYGQ 120 TARLNLPNIK NRGQLQGILL EEYLGLRNSD SSTTTSACSE STTTTSNQSE VCVPEEFTMR 180 PRLVSLNVKT EDGEGESKTC DHGDETATPM NQVKHEDRND QLVALGAEFP CLDQLENFQM 240 DEMFEPRTGT QAGHMVMPVS LEKQVKDEDL DAVYCGRSDD QAVLSEAGVP SLYDLHNFQM 300 DELFDVEELL SLINSDSLHD PTNIVKGNAD AYTNMAPSHV GSVGSEKPPN RSYQIQNPDA 360 KLLGSPQQME RTLADVDYGF DFLKQGREED LNAAADDCVR YLNEIGDLGF * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-17 | 70 | 127 | 2 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| 2gcc_A | 1e-17 | 70 | 134 | 5 | 70 | ATERF1 |
| 3gcc_A | 1e-17 | 70 | 134 | 5 | 70 | ATERF1 |
| 5wx9_A | 7e-17 | 59 | 133 | 3 | 78 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 9 | 16 | RKSRSRRD |
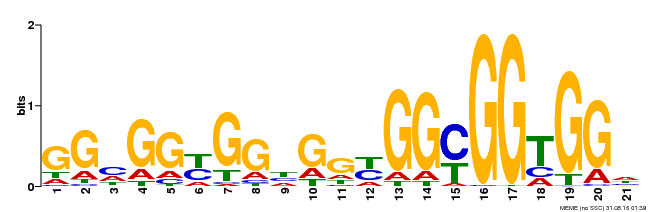
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00302 | DAP | Transfer from AT2G40340 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681815 | 0.0 | LN681815.1 Cucumis melo genomic scaffold, anchoredscaffold00008. | |||
| GenBank | LN713257 | 0.0 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004138558.1 | 0.0 | PREDICTED: dehydration-responsive element-binding protein 2C isoform X2 | ||||
| TrEMBL | A0A0A0KBP3 | 0.0 | A0A0A0KBP3_CUCSA; Dehydration-responsive element-binding protein 2C | ||||
| STRING | XP_004138558.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1175 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40340.1 | 3e-37 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.136780.1 |
| Entrez Gene | 101215431 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



