 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g002144m | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 961aa MW: 106084 Da PI: 5.2277 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92 | 4.3e-29 | 553 | 603 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
orange1.1g002144m 553 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 603
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.484 | 542 | 623 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.6E-26 | 556 | 603 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 4.25E-23 | 859 | 948 | No hit | No description |
| Gene3D | G3DSA:3.10.20.240 | 2.3E-27 | 864 | 945 | No hit | No description |
| PROSITE profile | PS51745 | 25.265 | 864 | 946 | IPR000270 | PB1 domain |
| SMART | SM00666 | 9.1E-26 | 864 | 946 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 8.4E-19 | 865 | 945 | IPR000270 | PB1 domain |
| CDD | cd06407 | 5.84E-39 | 865 | 945 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 961 aa Download sequence Send to blast |
MPEPDEENQT RYMPPAPPRT KEVTFMASSI MDVDVVDLDL DNPWPSDQMG FVSNPMSPFL 60 ISEQPCSPLW AFSDADNDDK LSGHNPDGYC MIKERITQAL RYFKDSTEQH VLAQVWVPVK 120 VGGRYVLTTS GQPFVLDPHS NGLHQYRMVS LMYMFSVDGE SDGELGLPGR VFWQKLPEWT 180 PNVQYYSSKE YSRLDHALHH NVRGTMALPV FEPSGQSCVA VIELIMTSQK INYAPEVDKV 240 CKALEAVNLK SSEILDYPST QICNEGRQNA LAEILEILSV VCETHKLPLA QTWVPCRHRS 300 VLAYGGGLKK SCSSIDGSCM GQVCMSTTDV AFYVVDGHMW GFREACVEHH LQKDQGVAGR 360 AFFSLSSCFC KDITQFCKTE YPLVHYARMF GLTSCFAICL RSTYTGDDDY ILEFFLPPAI 420 TDSYEQQTLL GSILATMKQH FQSLKVASGI DLEDDEGTIE IIEGEATADK KLNLRMESIR 480 IPQSVRSPPQ PHALPNGGEL GQLDIPEQQL MENFDYMNSR GNAVNVGGND NPVSLLENKN 540 TRKLSERKRG KTEKSISLEV LQQYFAGSLK DAAKSLGVCP TTMKRICRQH GISRWPSRKI 600 NKVNRSLTKL KRVIESVQGT NGTFGLTSLT TSPLPVAVNS ISWPSGLNGS NQQNSPNSKP 660 ELLGEKILSP IYKTPGSDGH TELEDRLSGG RMSTHEEHIH EQNALSPEIG KGKNSPKTGS 720 GSREESDGSP TSHGSCQGNP ANESAPAKDV LVSSIHEPRF KVGGSLELVF QPVKEMNLSA 780 AFSIPDALVT TEPQEPFGGL LVEDAGSSKD LRNLCPAVAD AIVDERLPEN SCANLPCAEL 840 SPKQHLATLS QTMPRVYSRQ EMKSVTIKAT YREDIIRFRI SLSCGILELK EEVAKRLKLE 900 LGTFDIKYLD DDQEWVLIAC DADLQECLDI SRSSGSNMIR LSIHDIMANL GSSCESTGEL 960 * |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
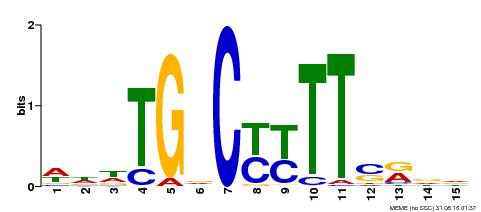
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KX056270 | 0.0 | KX056270.1 Citrus trifoliata transcription factor NLP7 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006476342.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A2H5NHG8 | 0.0 | A0A2H5NHG8_CITUN; Uncharacterized protein (Fragment) | ||||
| STRING | XP_006476342.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g002144m |



