 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna08492.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 1037aa MW: 114732 Da PI: 6.3272 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 76.7 | 2.6e-24 | 277 | 378 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-E CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDf 79
f+k+lt sd++++g +++p++ ae+ ++++ + ++l+++d++ ++W++++iyr++++r++lt+GW+ Fv ++Lk+gD+
mrna08492.1-v1.0-hybrid 277 FCKTLTASDTSTHGGFSVPRRAAEKLfppldftMQPP-T-QELVVRDLHDNTWTFRHIYRGQPKRHLLTTGWSLFVGTKRLKAGDS 360
99*********************999*****954444.4.48******************************************** PP
EEEEE-SSSEE..EEEEE-S CS
B3 80 vvFkldgrsefelvvkvfrk 99
v+F ++++ +l+v+++r+
mrna08492.1-v1.0-hybrid 361 VLFI--RDEKSQLLVGIRRA 378
****..4577778*****97 PP
| |||||||
| 2 | Auxin_resp | 115.6 | 4.5e-38 | 403 | 486 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalk.vkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaa+++s+F+++YnPra++seFv+++ k++kal+ +++svGmRf m+fete+ss+rr++Gt+v +sdldp+rWp+SkWr+L+
mrna08492.1-v1.0-hybrid 403 AAHAAANRSPFTIFYNPRACPSEFVIPLVKFQKALYgNQLSVGMRFGMMFETEESSKRRYMGTIVNISDLDPLRWPGSKWRNLQ 486
79**********************************9*********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 2.62E-46 | 264 | 406 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 8.5E-42 | 270 | 391 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.54E-23 | 276 | 377 | No hit | No description |
| Pfam | PF02362 | 1.7E-22 | 277 | 378 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.7E-25 | 277 | 379 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.14 | 277 | 379 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.2E-33 | 403 | 486 | IPR010525 | Auxin response factor |
| SuperFamily | SSF54277 | 6.38E-7 | 933 | 1005 | No hit | No description |
| PROSITE profile | PS51745 | 18.598 | 936 | 1026 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0007134 | developmental stage | sporophyte vegetative stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1037 aa Download sequence Send to blast |
MEETCETEKR RRDGPDRIPT RVNPGNVRLP VLTFVVNGTR GRSKWGWWFI ERSGSFQEVK 60 TSTRLKTHAL VGVPFDAKLV MGRRAKGRRG GGGEPELEIG FYRILRNRVS FKFWKEYGTK 120 LSLVRVRGER VIMEEKMRVG GGLLSGAQSS ILEEMKLLKE LQDHSGSRKA INSELWHACA 180 GPLVCLPQVG SLAYYFPQGH SEQVAVSTKR IATSQIPNYP NLPSQLLCQV QNVTLHADKE 240 TDEIFTQMCL KPVNSEKDVF PVPDFGLKPS KHPSEFFCKT LTASDTSTHG GFSVPRRAAE 300 KLFPPLDFTM QPPTQELVVR DLHDNTWTFR HIYRGQPKRH LLTTGWSLFV GTKRLKAGDS 360 VLFIRDEKSQ LLVGIRRANR QQTTLPSSVL SADSMHIGVL AEAAHAAANR SPFTIFYNPR 420 ACPSEFVIPL VKFQKALYGN QLSVGMRFGM MFETEESSKR RYMGTIVNIS DLDPLRWPGS 480 KWRNLQVEWD EPGCCDKQNR VSSWEVETPE SLFIFPSLTS SLKRPFHPGY LSAETEWANM 540 IKRPFIRVPE MGHMNSLPYQ MSNLCSEQLV NMLLKPQLVS QAGTLSALQQ ESAANGGALE 600 DMQAMQAKMN QKNLAFCSEG MSLQSQNPSQ SCTTAKFGSQ TPVGANTDKT KLEPDLSTDQ 660 VSQLSSTGQG NEEKLAAGIA SSPYNHAFVN QNQGQLQTSP RPMQQPMESL LYHSQQTDLP 720 QSDFNSANSS LPSIENDECM FYQPFAGILR SPGPLSAYGL QDSPSVLTEA NNFSLTSVGQ 780 EMWDNSLSRL LPQVDQLTSS HQDLSTFNSI PNSGSLRDLS DESNNQSGVY GCPSVDVGTG 840 VANIVADPSV TSTIMDEFSK LKHAEFHNPS ECLVGNLSSS QDLQSQITSA SLGDSQAFSR 900 QELADNSGGT SSSNVDLDES SLLQNNSWHQ VVPPVRTYTK VQKAGSVGRS IDVTSYTNYE 960 ELCSAIECMF GLEGLLNDPR GSGWKLVYVD YENDVLLVGD DPWEILSPKE VQEMSEEGMK 1020 LLNSAAALQG MHTTVS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 147 | 511 | 29 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
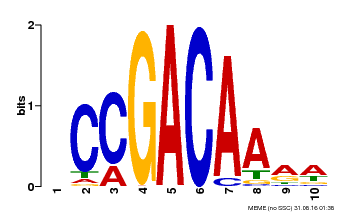
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna08492.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004291385.1 | 0.0 | PREDICTED: auxin response factor 5 | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| TrEMBL | A0A2P6PZ09 | 0.0 | A0A2P6PZ09_ROSCH; Auxin response factor | ||||
| STRING | XP_004291385.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8991 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna08492.1-v1.0-hybrid |



