 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010910785.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 282aa MW: 29651.3 Da PI: 7.127 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.9 | 0.00071 | 129 | 151 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y+C+ C+k+F++ L H +H
XP_010910785.1 129 YECKTCNKCFPSFQALGGHRASH 151
99**********99999998887 PP
| |||||||
| 2 | zf-C2H2 | 13.3 | 0.00025 | 195 | 217 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++Cg F++ L H+r+H
XP_010910785.1 195 HECSICGSEFNSGQALGGHMRRH 217
78********************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.36E-9 | 128 | 151 | No hit | No description |
| Pfam | PF13912 | 9.3E-12 | 128 | 153 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 10.221 | 129 | 156 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.082 | 129 | 151 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 131 | 151 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.36E-9 | 190 | 217 | No hit | No description |
| Pfam | PF13912 | 5.2E-12 | 194 | 219 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.598 | 195 | 217 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.021 | 195 | 217 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 197 | 217 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 282 aa Download sequence Send to blast |
MESPEEAMGS NSTTSNGNGG ECTHATSTAI VVKGKRTKRP RTQPLALAAA MAVADSSSAS 60 SADASGSITE EEEDMANCLI LLAQGRALDA GGPKPELPAE EGGGTDKFTS RRFTEAATTT 120 GGKAGFYVYE CKTCNKCFPS FQALGGHRAS HKKPKLAITA GEEKKASVEE DMLKISMNSF 180 SKAMVSGGST KPRVHECSIC GSEFNSGQAL GGHMRRHRPV VIPEAAEAKK ERTILSLDLN 240 LPAPSDDDRP ELQRPSSPAF SFASKPPLIF PASASALVDC HY |
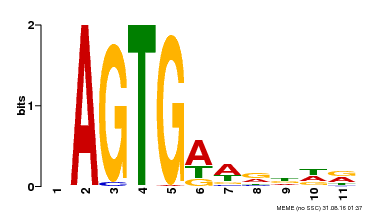
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00487 | DAP | Transfer from AT5G04390 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010910785.1 | 0.0 | zinc finger protein ZAT5 | ||||
| TrEMBL | A0A2H3Z3L2 | 1e-162 | A0A2H3Z3L2_PHODC; zinc finger protein ZAT5 | ||||
| STRING | XP_008807698.1 | 1e-163 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2013 | 32 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G04390.1 | 2e-50 | C2H2 family protein | ||||



