 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00012698-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 995aa MW: 109636 Da PI: 4.6846 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92 | 4.5e-29 | 603 | 653 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
WALNUT_00012698-RA 603 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 653
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.564 | 592 | 673 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.8E-26 | 606 | 653 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.17E-22 | 897 | 986 | No hit | No description |
| PROSITE profile | PS51745 | 24.519 | 900 | 982 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 4.3E-26 | 900 | 982 | No hit | No description |
| SMART | SM00666 | 5.8E-27 | 900 | 982 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 2.5E-19 | 901 | 981 | IPR000270 | PB1 domain |
| CDD | cd06407 | 7.22E-37 | 901 | 981 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 995 aa Download sequence Send to blast |
MCGPEELNSC GLPPKSTGPQ VVEEEEEEEE QEEEEEEAAA EEERVVEALI MDFDDLDGSW 60 PLDQVSSVSD HNDDDKNNNQ YPMSPLYLSC SDQAFSPLWA FLDGNDDASH ATSAPPVCSA 120 LFPGTTIPVA ERPVVNNFDD SRLPFPLMGL TPQENPDGCF VIKERMTLAL RYFKDLTEQN 180 VLAQVWVPVK NGSRYVLTTS GQPFVLDPHS NGLNQYRMVS QTYVFPVDED NNGLLGLPGR 240 VFCQRVPEWT PNVQYYSMLE YPRRNHAQNF NVRGSLALPV FEPSGESCVG VLELILTSAK 300 INYAPEVDKV CKALEICNEG RQNALAEILG ILTVVCETHK LPLAQTWVPC MHRNILAYGG 360 GLKKSCTSFD GSCMAQICMS TPDVALYVVD SQMWRFREAC VQHHLKKGQG VAGRAFLSRS 420 SCFCGNITLF CKTNYPLVHY ARMFGLAGCF AICLRSTHTG DDDYILEFFL PPGITDFYEQ 480 QILLGSLLAT MKQHFQSLKV ASDIELEEEV SVEIVQVSED GVDSRIESIR IPLSTKSTPR 540 PDGLLDMGDM VQDSPKQQLM VHLDDINDKG TDVKNGGGSI NNFSSLDIKE MKNKSQRKRG 600 KTEKSISLEV LQQYFAGSLK DAAKSLGVCP TTMKRICRQH GISRWPSRKI NKVNRSLTKL 660 KRVIESVQGA EGTFSLNSLN PSPLPIRSNQ QTSPCFKHSE PHEKNESPTR ITLKSDEQDG 720 MEDLLQGGRT FSPEKLVHDH SRSPPEVGKG SNGSKARSGS CEESAGTPTS HGSCQGGPPV 780 ESTALAKHLY VSSINDQCVK AGGDSPELAY PPTGPPNISA AYSLPDGILM TEPQETFGGM 840 LIEDAGSSKD LRNLCPSVAD AVLDEQAPEF CLPNPPCPNQ SMPTLAPTTP HVTARKEMKN 900 VTIKATYKED IIRFRISLSS GILELKEEVA KRLKLEVGTF DIKYLDDDLE WVLIACDADL 960 QECMDIPRSS GSNIIRLLVQ DTISNLGSSC ESSWD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
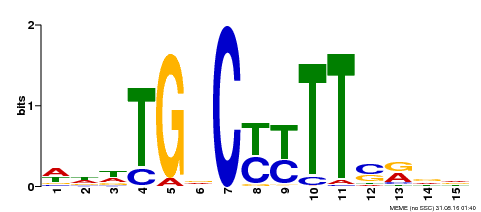
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018824315.1 | 0.0 | PREDICTED: protein NLP6-like | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A2I4EY38 | 0.0 | A0A2I4EY38_JUGRE; protein NLP6-like | ||||
| STRING | EOY25090 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4257 | 34 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



