 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi11g11880.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 665aa MW: 72058.8 Da PI: 6.3088 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 79.3 | 4.8e-25 | 286 | 336 | 1 | 52 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQk 52
kpr++W+ eLH++Fv+av+qL G +kA+Pk+ilelm+v+gLt+e+v+SHLQ
Vradi11g11880.1 286 KPRVVWSVELHQQFVSAVNQL-GLDKAVPKRILELMNVPGLTRENVASHLQG 336
79*******************.*****************************6 PP
| |||||||
| 2 | Response_reg | 74.7 | 3.5e-25 | 104 | 212 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealka 92
vl+vdD+ + +++++q+ + +y +v+++++++ al+ll+e++ +D++l D+ mp+mdG++ll++ e +lp+i++++ ++ + + + ++
Vradi11g11880.1 104 VLVVDDDATTLRIIEQMSIRCRY-RVTTCTEATVALNLLRERKgcFDVVLSDVHMPDMDGYKLLEHVGLEM-DLPVIMMSGDSTTSAVMKGIRH 195
89*********************.*******************************************6644.8********************* PP
TESEEEESS--HHHHHH CS
Response_reg 93 GakdflsKpfdpeelvk 109
Ga d+l Kp+ +eel++
Vradi11g11880.1 196 GACDYLIKPVREEELRN 212
***************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 2.8E-39 | 101 | 233 | No hit | No description |
| SuperFamily | SSF52172 | 1.31E-33 | 101 | 225 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 4.9E-29 | 102 | 214 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 40.387 | 103 | 218 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.6E-21 | 104 | 213 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.13E-25 | 105 | 217 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-25 | 283 | 336 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 9.799 | 283 | 342 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.94E-16 | 283 | 340 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-21 | 286 | 336 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.9E-7 | 288 | 338 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 665 aa Download sequence Send to blast |
MDTITALAGT MLLVGSGELH GRFLCSQITL LALKQPNEGV GDDAFKIRVS ILLLCSLSAI 60 VLRFSASELS PDHVVALVAL MAQSRSAGDA CKGGTAEFPA GLRVLVVDDD ATTLRIIEQM 120 SIRCRYRVTT CTEATVALNL LRERKGCFDV VLSDVHMPDM DGYKLLEHVG LEMDLPVIMM 180 SGDSTTSAVM KGIRHGACDY LIKPVREEEL RNIWQHVVRK IWNENKEHDN SGSMEDSDWN 240 KRGNDDTEYT SSVADAAEVV KAPKKRSSLK EEDIELESDD PATSKKPRVV WSVELHQQFV 300 SAVNQLGLDK AVPKRILELM NVPGLTRENV ASHLQGWLMR LPVVNEKFRL YLKRLSGVAQ 360 QQNGMLNAVP GTIESKLSAT GRFDIQALAA AGHVPPETLA ALHAELLGRP ATSIMSTADQ 420 ATLLQASIGL KHSHAEHAVA YGQPFVKCPS NIGKDFPPSF LTANDVSSGY GAWPSSNNTM 480 GSAVNGSASV NQNCSFGMNA IIDYSLLSQK SNNSVSASQF SGGDTKTQAV LYAHTTPSTC 540 SMSSINGSIP QLQNSTLTFG GARPLPSPLP MSDRQDPYGL KSGDSFDQVC LRNIGFIGKG 600 TCIPNRFVQN EIESSVSDFS QMKVNVDSTG NTLKEEPNFI NMQKVSRPIF GRYQSCDHSS 660 VFTE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-19 | 285 | 356 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
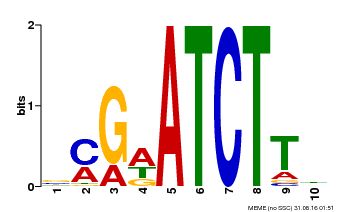
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi11g11880.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015043 | 0.0 | AP015043.1 Vigna angularis var. angularis DNA, chromosome 10, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022643111.1 | 0.0 | two-component response regulator ORR21 isoform X2 | ||||
| Swissprot | A2XE31 | 1e-141 | ORR21_ORYSI; Two-component response regulator ORR21 | ||||
| Swissprot | Q8H7S7 | 1e-141 | ORR21_ORYSJ; Two-component response regulator ORR21 | ||||
| TrEMBL | A0A3Q0FHC8 | 0.0 | A0A3Q0FHC8_VIGRR; Two-component response regulator | ||||
| STRING | GLYMA02G09451.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF11808 | 24 | 33 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-112 | response regulator 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



