 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA4973 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 418aa MW: 44944.8 Da PI: 8.3931 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 63.5 | 4e-20 | 271 | 333 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkrerrkq+NRe+ArrsR+RK+ae eeL++kv++L ae +Lk+e+++l + ++kl++e+
Tp57577_TGAC_v2_mRNA4973 271 ERELKRERRKQSNRESARRSRLRKQAEAEELARKVESLNAESASLKSEINRLAESSEKLRMEN 333
89**********************************************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 1.9E-36 | 1 | 98 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 3.8E-24 | 131 | 252 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 3.0E-16 | 265 | 331 | No hit | No description |
| Pfam | PF00170 | 1.8E-18 | 271 | 333 | IPR004827 | Basic-leucine zipper domain |
| SMART | SM00338 | 1.9E-18 | 271 | 335 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.162 | 273 | 336 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.88E-9 | 274 | 330 | No hit | No description |
| CDD | cd14702 | 1.04E-19 | 276 | 324 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 278 | 293 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 418 aa Download sequence Send to blast |
MGTSEEEKST KTEKPSSPLT ADQTNNQANI HVYPDWAAMQ AYYGPRVAMP PYYNSPVPSG 60 HTPHPYMWGP PQPMMPPYGH PYAAMYPHGG VYTHPAVPIV THPHSQGISS SPVTGTPLSI 120 ETPPKSSGNT DQGLMKKLKG FDGLAMSIGN GHAESAEPGA ESKLSQSVDT DGSSDGSDSN 180 TSGANQTRRK RSREGTPTTD DGEGKTHTQG SQASKEIATS NKMMAVGPVV SSAMTTALEL 240 KNPSSVKSKT SPTSAPQPSA VLPPEAWMQN ERELKRERRK QSNRESARRS RLRKQAEAEE 300 LARKVESLNA ESASLKSEIN RLAESSEKLR MENAALKEKF KIAQLGQPKE IILSSIDSQR 360 ITPVSTENLL SRVNNNSGSN DRAMEDENSY CDNKPNSGAK LHQLLDTSPR ADAVAAG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 287 | 293 | RRSRLRK |
| 2 | 287 | 294 | RRSRLRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the G-box-like motif (5'-ACGTGGC-3') of the chalcone synthase (CHS) gene promoter. G-box and G-box-like motifs are defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
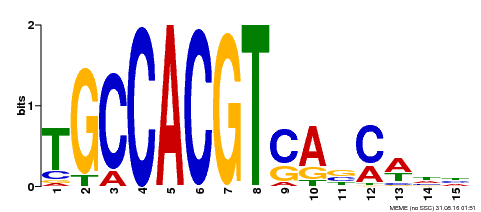
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA4973 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN863911 | 0.0 | JN863911.1 Medicago sativa bZIP protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024626445.1 | 0.0 | common plant regulatory factor 1 isoform X1 | ||||
| Refseq | XP_024626446.1 | 0.0 | common plant regulatory factor 1 isoform X1 | ||||
| Refseq | XP_024626447.1 | 0.0 | common plant regulatory factor 1 isoform X1 | ||||
| Refseq | XP_024626448.1 | 0.0 | common plant regulatory factor 1 isoform X1 | ||||
| Swissprot | Q99089 | 1e-137 | CPRF1_PETCR; Common plant regulatory factor 1 | ||||
| TrEMBL | A0A2K3PK46 | 0.0 | A0A2K3PK46_TRIPR; Common plant regulatory factor 1-like protein (Fragment) | ||||
| STRING | AES82747 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5301 | 34 | 55 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 3e-97 | G-box binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA4973 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



