 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10024328m | ||||||||
| Common Name | EUTSA_v10024328mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 959aa MW: 105598 Da PI: 5.6812 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.9 | 4.9e-29 | 587 | 637 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
Thhalv10024328m 587 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 637
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.757 | 576 | 657 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.3E-26 | 589 | 637 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.6E-23 | 859 | 947 | No hit | No description |
| Pfam | PF00564 | 1.9E-20 | 862 | 943 | IPR000270 | PB1 domain |
| SMART | SM00666 | 1.6E-28 | 862 | 944 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 27.83 | 862 | 944 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.7E-27 | 862 | 943 | No hit | No description |
| CDD | cd06407 | 1.51E-34 | 863 | 943 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 959 aa Download sequence Send to blast |
MCEPDDNTAR NGVTPQASRS RELLMDVDDL DLDGSWPLDQ IPYLSSASRM ISPIFVSSSS 60 EQPCSPLWAF SDGGGTNHAT AGGDDEKISS ASGVPSFRLA DYPLFLPYSS PSVAENTAEK 120 QNSFQFPSPL MSLVPPENAD NYCVIKERMT QALRYFKEST EQHVLAQVWA PVRKNGRDLL 180 TTLGQPFVLN PNGNGLNQYR MISLTYMFSV DSESDIELGL PGRVFRQKLP EWTPNVQYYS 240 SKEFSRLDHA LHYNVRGTLA LPVFNLSGES CIGVVELIMT SEKIHYAPEV DRVCKALEAV 300 NLKSSEILDH QTTQICNESR QNALAEILEV LTVVCETHNL PLAQTWVPCQ HGSVLANGGG 360 LKKNCTSFDG SCMGQVCMST TDMACYVVDA HVWGFRDACL EHHLQKGQGV AGRAFLNGGS 420 CFCRDITKFC KTQYPLVHYA LMFKLTTCFA ISLQSSYTGD DSYILEFFLP SSITDDQEQD 480 ALLGSLLVTM KEHFQSLRVA SGVDFGEDDD KLSFEIIRAL PDKKIHSKIE SIRVPFSGFK 540 SNATETLLIP QHAVQSSDPV NEKSNVATVN GVVKEKKKTE KKRGKTEKTI SLDVLQQYFT 600 GSLKDAAKSL GVCPTTMKRI CRQHGISRWP SRKIKKVNRS ITKLKRVIES VQGTDGGLDL 660 TSMAVSSIPW THGHQTAAQP LNSPSGSKPP ELPNTDNSPN HWSSDHSPHE PNGSPELPNN 720 GHKRSRTADE SAGTPTSHGS CDGNQLDETK VLNQDPLFTV GGSPGLFFPP YSRDHDVSAA 780 SFSLPNRLLG TIDHFRGMPI EDAGSSKDLR NLCSTAAFED KFPDSNWMTN DNNSNNNNIY 840 APPKEEAIVN ITREPSGSEM RTVTIKASYK EDIIRFRISS GSGIMELKDE VAKRLKLDAG 900 TFDIKYLDDD NEWVLIACDA DLQECLEIPR SSRTNIVRLL VHDATTNLGS SCESTGEL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
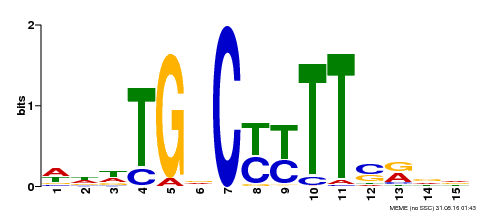
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10024328m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228452 | 0.0 | AK228452.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-03-P13. | |||
| GenBank | BT005776 | 0.0 | BT005776.1 Arabidopsis thaliana clone RAFL15-03-P13 (R20174) unknown protein (At4g24020) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006413484.1 | 0.0 | protein NLP7 | ||||
| Refseq | XP_024005488.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | V4MHY8 | 0.0 | V4MHY8_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006413484.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10024328m |
| Entrez Gene | 18030707 |



